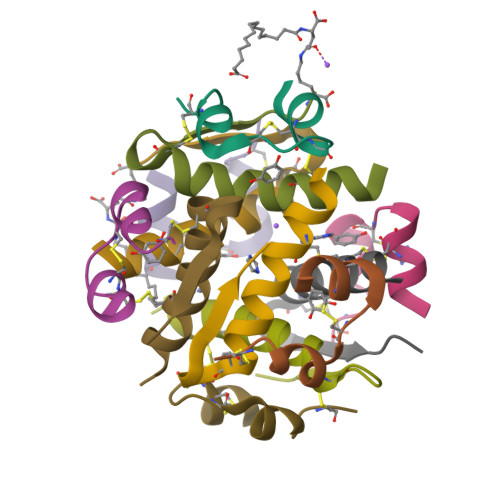
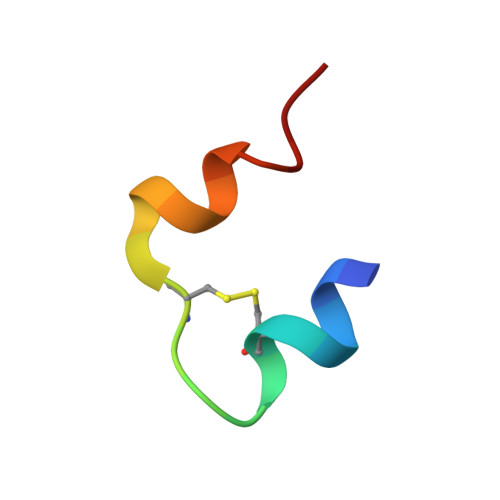Ligand Controlled Assembly of Hexamers, Dihexamers, and Linear Multihexamer Structures by the Engineered Acylated Insulin Degludec.
Steensgaard, D.B., Schluckebier, G., Strauss, H.M., Norrman, M., Thomsen, J.K., Friderichsen, A.V., Havelund, S., Jonassen, I.(2013) Biochemistry 52: 295
- PubMed: 23256685
- DOI: https://doi.org/10.1021/bi3008609
- Primary Citation of Related Structures:
4AJX, 4AJZ, 4AK0, 4AKJ - PubMed Abstract:
Insulin degludec, an engineered acylated insulin, was recently reported to form a soluble depot after subcutaneous injection with a subsequent slow release of insulin and an ultralong glucose-lowering effect in excess of 40 h in humans. We describe the structure, ligand binding properties, and self-assemblies of insulin degludec using orthogonal structural methods. The protein fold adopted by insulin degludec is very similar to that of human insulin. Hexamers in the R(6) state similar to those of human insulin are observed for insulin degludec in the presence of zinc and resorcinol. However, under conditions comparable to the pharmaceutical formulation comprising zinc and phenol, insulin degludec forms finite dihexamers that are composed of hexamers in the T(3)R(3) state that interact to form an R(3)T(3)-T(3)R(3) structure. When the phenolic ligand is depleted and the solvent condition thereby mimics that of the injection site, the quaternary structure changes from dihexamers to a supramolecular structure composed of linear arrays of hundreds of hexamers in the T(6) state and an average molar mass, M(0), of 59.7 × 10(3) kg/mol. This novel concept of self-assemblies of insulin controlled by zinc and phenol provides the basis for the slow action profile of insulin degludec. To the best of our knowledge, this report for the first time describes a tight linkage between quaternary insulin structures of hexamers, dihexamers, and multihexamers and their allosteric state and its origin in the inherent propensity of the insulin hexamer for allosteric half-site reactivity.
Organizational Affiliation:
Diabetes Protein Engineering, Novo Nordisk A/S , Novo Nordisk Park, 2760 Maaloev, Denmark. dbs@novonordisk.com