Crystal Structure of Metallo-beta-Lactamase SMB-1 Bound to Hydrolyzed Imipenem
Wachino, J., Arakawa, Y.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Metallo-beta-lactamase | 262 | Serratia marcescens | Mutation(s): 0 Gene Names: SMB-1 EC: 3.5.2.6 | 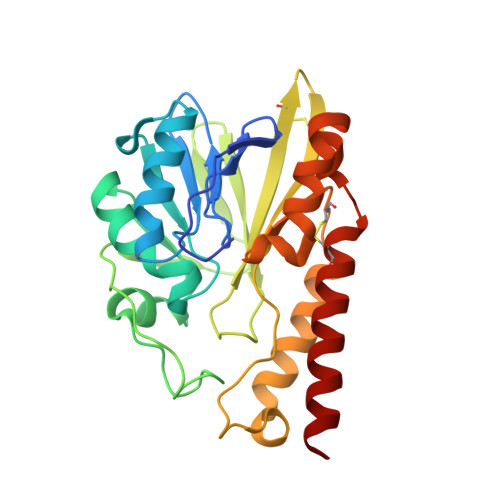 | |
UniProt | |||||
Find proteins for G5ELM3 (Serratia marcescens) Explore G5ELM3 Go to UniProtKB: G5ELM3 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | G5ELM3 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| HIW Query on HIW | V [auth A] | (2R,4S)-2-[(1S,2R)-1-carboxy-2-hydroxypropyl]-4-[(2-{[(Z)-iminomethyl]amino}ethyl)sulfanyl]-3,4-dihydro-2H-pyrrole-5-ca
rboxylic acid C12 H19 N3 O5 S GGEWNUMDSNUHAH-LURQLKTLSA-N |  | ||
| SO4 Query on SO4 | S [auth A], T [auth A], U [auth A] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| ZN Query on ZN | B [auth A], C [auth A] | ZINC ION Zn PTFCDOFLOPIGGS-UHFFFAOYSA-N |  | ||
| NA Query on NA | D [auth A] E [auth A] F [auth A] G [auth A] H [auth A] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 37.03 | α = 108.32 |
| b = 41.46 | β = 102.78 |
| c = 45.44 | γ = 106.28 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| iMOSFLM | data reduction |
| SCALA | data scaling |
| MOLREP | phasing |