Observing cellulose biosynthesis and membrane translocation in crystallo.
Morgan, J.L., McNamara, J.T., Fischer, M., Rich, J., Chen, H.M., Withers, S.G., Zimmer, J.(2016) Nature 531: 329-334
- PubMed: 26958837
- DOI: https://doi.org/10.1038/nature16966
- Primary Citation of Related Structures:
5EIY, 5EJ1, 5EJZ - PubMed Abstract:
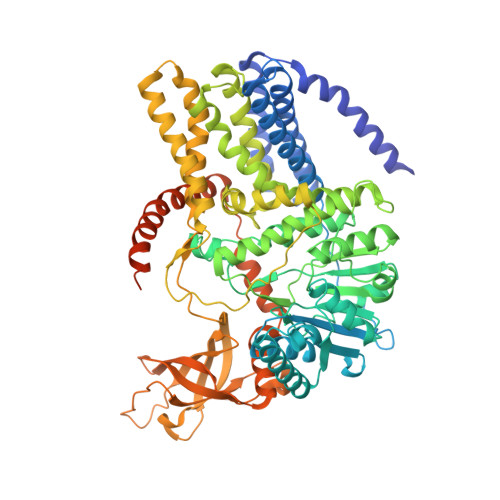
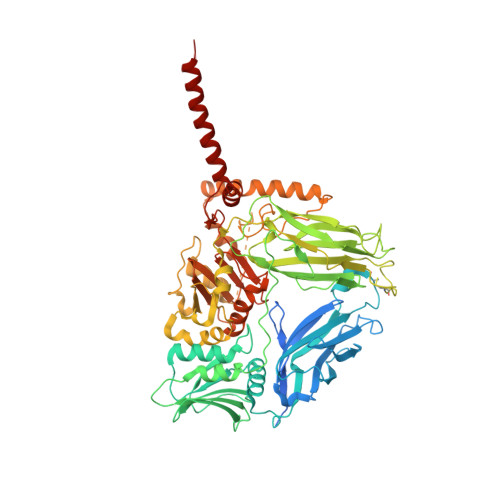
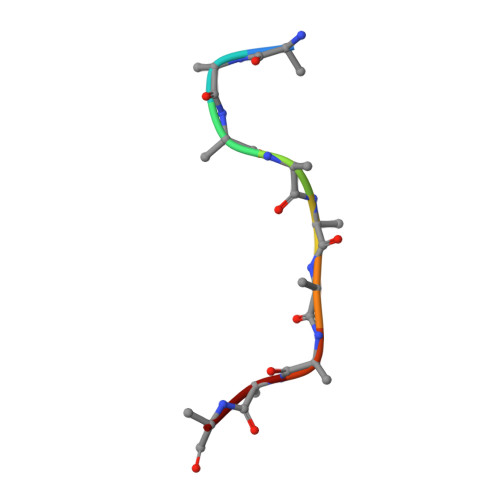
Many biopolymers, including polysaccharides, must be translocated across at least one membrane to reach their site of biological function. Cellulose is a linear glucose polymer synthesized and secreted by a membrane-integrated cellulose synthase. Here, in crystallo enzymology with the catalytically active bacterial cellulose synthase BcsA-BcsB complex reveals structural snapshots of a complete cellulose biosynthesis cycle, from substrate binding to polymer translocation. Substrate- and product-bound structures of BcsA provide the basis for substrate recognition and demonstrate the stepwise elongation of cellulose. Furthermore, the structural snapshots show that BcsA translocates cellulose via a ratcheting mechanism involving a 'finger helix' that contacts the polymer's terminal glucose. Cooperating with BcsA's gating loop, the finger helix moves 'up' and 'down' in response to substrate binding and polymer elongation, respectively, thereby pushing the elongated polymer into BcsA's transmembrane channel. This mechanism is validated experimentally by tethering BcsA's finger helix, which inhibits polymer translocation but not elongation.
Organizational Affiliation:
University of Virginia School of Medicine, Center for Membrane Biology, Molecular Physiology and Biological Physics, 480 Ray C. Hunt Drive, Charlottesville, Virginia 22908, USA.