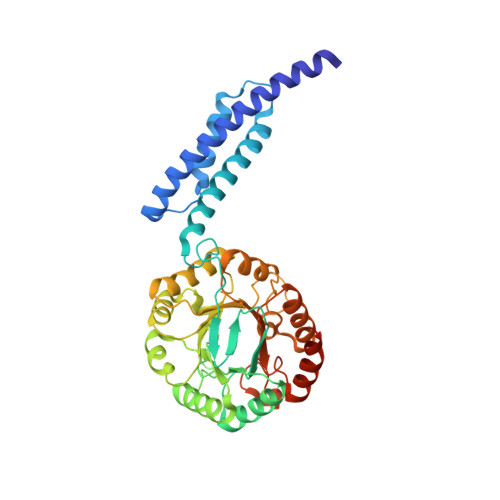
Interdomain Conformational Changes Provide Allosteric Regulation en Route to Chorismate.
Nazmi, A.R., Lang, E.J.M., Bai, Y., Allison, T.M., Othman, M.H., Panjikar, S., Arcus, V.L., Parker, E.J.(2016) J Biol Chem 291: 21836-21847
- PubMed: 27502275
- DOI: https://doi.org/10.1074/jbc.M116.741637
- Primary Citation of Related Structures:
5J6F - PubMed Abstract:
Multifunctional proteins play a variety of roles in metabolism. Here, we examine the catalytic function of the combined 3-deoxy-d- arabino heptulosonate-7-phosphate synthase (DAH7PS) and chorismate mutase (CM) from Geobacillus sp. DAH7PS operates at the start of the biosynthetic pathway for aromatic metabolites, whereas CM operates in a dedicated branch of the pathway for the biosynthesis of amino acids tyrosine and phenylalanine. In line with sequence predictions, the two catalytic functions are located in distinct domains, and these two activities can be separated and retain functionality. For the full-length protein, prephenate, the product of the CM reaction, acts as an allosteric inhibitor for the DAH7PS. The crystal structure of the full-length protein with prephenate bound and the accompanying small angle x-ray scattering data reveal the molecular mechanism of the allostery. Prephenate binding results in the tighter association between the dimeric CM domains and the tetrameric DAH7PS, occluding the active site and therefore disrupting DAH7PS function. Acquisition of a physical gating mechanism to control catalytic function through gene fusion appears to be a general mechanism for providing allostery for this enzyme.
Organizational Affiliation:
From the Biomolecular Interaction Centre and Department of Chemistry, University of Canterbury, P. O. Box 4800, Christchurch 8140, New Zealand.