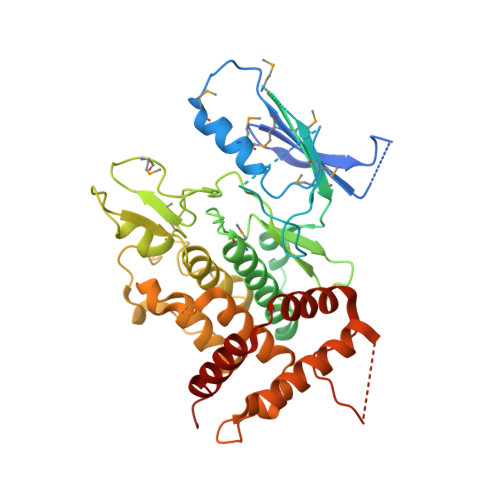
Structural insights into ubiquitin phosphorylation by PINK1.
Okatsu, K., Sato, Y., Yamano, K., Matsuda, N., Negishi, L., Takahashi, A., Yamagata, A., Goto-Ito, S., Mishima, M., Ito, Y., Oka, T., Tanaka, K., Fukai, S.(2018) Sci Rep 8: 10382-10382
- PubMed: 29991771
- DOI: https://doi.org/10.1038/s41598-018-28656-8
- Primary Citation of Related Structures:
5YJ9 - PubMed Abstract:
Mutations of PTEN-induced putative kinase 1 (PINK1) and the E3 ubiquitin (Ub) ligase parkin can cause familial parkinsonism. These two proteins are essential for ubiquitylation of damaged mitochondria and subsequent degradation. PINK1 phosphorylates Ser65 of Ub and the Ub-like (UBL) domain of parkin to allosterically relieve the autoinhibition of parkin. To understand the structural mechanism of the Ub/UBL-specific phosphorylation by PINK1, we determined the crystal structure of Tribolium castaneum PINK1 kinase domain (TcPINK1) in complex with a nonhydrolyzable ATP analogue at 2.5 Å resolution. TcPINK1 consists of the N- and C-terminal lobes with the PINK1-specific extension. The ATP analogue is bound in the cleft between the N- and C-terminal lobes. The adenine ring of the ATP analogue is bound to a hydrophobic pocket, whereas the triphosphate group of the ATP analogue and two coordinated Mg ions interact with the catalytic hydrophilic residues. Comparison with protein kinases A and C (PKA and PKC, respectively) unveils a putative Ub/UBL-binding groove, which is wider than the peptide-binding groove of PKA or PKC to accommodate the globular head of Ub or UBL. Further crosslinking analyses suggested a PINK1-interacting surface of Ub. Structure-guided mutational analyses support the findings from the present structural analysis of PINK1.
Organizational Affiliation:
Institute for Quantitative Biosciences, The University of Tokyo, Tokyo, 113-0032, Japan.