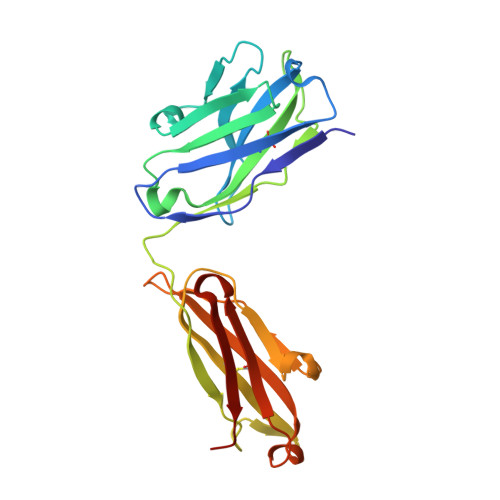
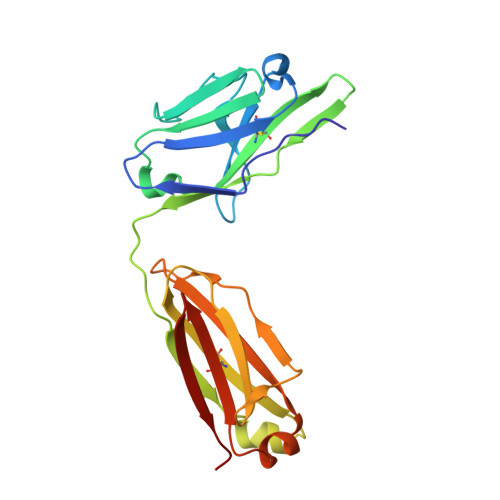
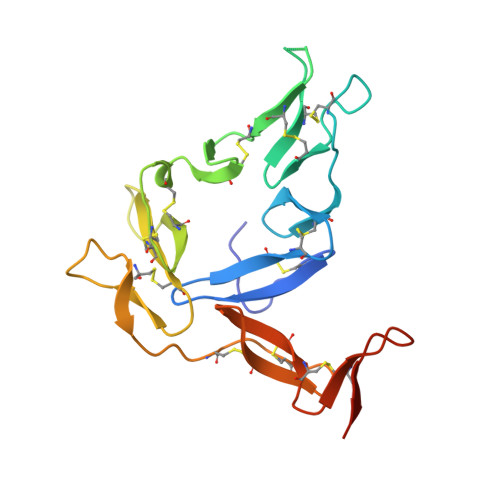
Molecular definition of multiple sites of antibody inhibition of malaria transmission-blocking vaccine antigen Pfs25.
Scally, S.W., McLeod, B., Bosch, A., Miura, K., Liang, Q., Carroll, S., Reponen, S., Nguyen, N., Giladi, E., Ramisch, S., Yusibov, V., Bradley, A., Lemiale, F., Schief, W.R., Emerling, D., Kellam, P., King, C.R., Julien, J.P.(2017) Nat Commun 8: 1568-1568
- PubMed: 29146922
- DOI: https://doi.org/10.1038/s41467-017-01924-3
- Primary Citation of Related Structures:
6AZZ, 6B08, 6B0A, 6B0E, 6B0G, 6B0H - PubMed Abstract:
The Plasmodium falciparum Pfs25 protein (Pfs25) is a leading malaria transmission-blocking vaccine antigen. Pfs25 vaccination is intended to elicit antibodies that inhibit parasite development when ingested by Anopheles mosquitoes during blood meals. The Pfs25 three-dimensional structure has remained elusive, hampering a molecular understanding of its function and limiting immunogen design. We report six crystal structures of Pfs25 in complex with antibodies elicited by immunization via Pfs25 virus-like particles in human immunoglobulin loci transgenic mice. Our structural findings reveal the fine specificities associated with two distinct immunogenic sites on Pfs25. Importantly, one of these sites broadly overlaps with the epitope of the well-known 4B7 mouse antibody, which can be targeted simultaneously by antibodies that target a non-overlapping site to additively increase parasite inhibition. Our molecular characterization of inhibitory antibodies informs on the natural disposition of Pfs25 on the surface of ookinetes and provides the structural blueprints to design next-generation immunogens.
Organizational Affiliation:
Program in Molecular Medicine, The Hospital for Sick Children Research Institute, 686 Bay St, Toronto, ON, Canada, M5G 0A4.