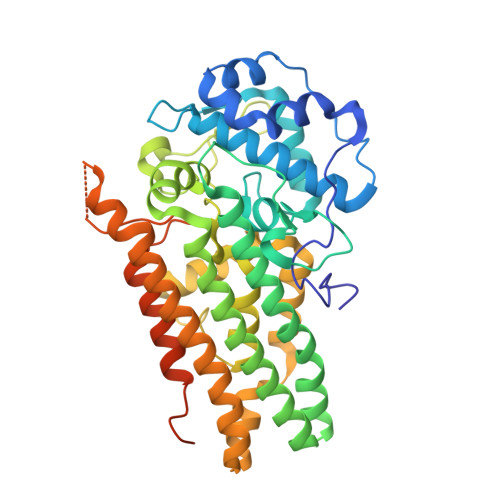
Inhibition Mechanisms of Human Indoleamine 2,3 Dioxygenase 1.
Lewis-Ballester, A., Karkashon, S., Batabyal, D., Poulos, T.L., Yeh, S.R.(2018) J Am Chem Soc 140: 8518-8525
- PubMed: 29897749
- DOI: https://doi.org/10.1021/jacs.8b03691
- Primary Citation of Related Structures:
6CXU, 6CXV - PubMed Abstract:
Human indoleamine 2,3-dioxygenase 1 (hIDO1) and tryptophan dioxygenase (hTDO) catalyze the same dioxygenation reaction of Trp to generate N-formyl kynurenine (NFK). They share high structural similarity, especially in the active site. However, hIDO1 possesses a unique inhibitory substrate binding site (Si) that is absent in hTDO. In addition, in hIDO1, the indoleamine group of the substrate Trp is H-bonded to S167 through a bridging water, while that in hTDO is directly H-bonded to H76. Here we show that Trp binding to the Si site or the mutation of S167 to histidine in hIDO1 retards its turnover activity and that the inhibited activity can be rescued by an effector, 3-indole ethanol (IDE). Kinetic studies reveal that the inhibited activity introduced by Trp binding to the Si site is a result of retarded recombination of the ferryl moiety with Trp epoxide to form NFK and that IDE reverses the effect by preventing Trp from binding to the Si site. In contrast, the abolished activity induced by the S167H mutation is primarily a result of ∼5000-fold reduction in the O 2 binding rate constant, possibly due to the blockage of a ligand delivery tunnel, and that IDE binding to the Si site reverses the effect by reopening the tunnel. The data offer new insights into structure-based design of hIDO1-selective inhibitors.
Organizational Affiliation:
Department of Physiology and Biophysics , Albert Einstein College of Medicine , Bronx , New York 10461 , United States.