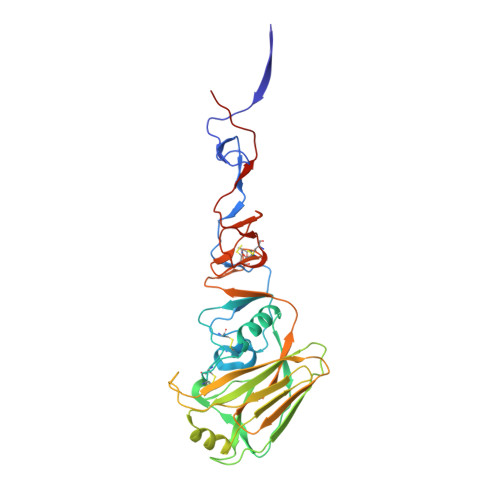
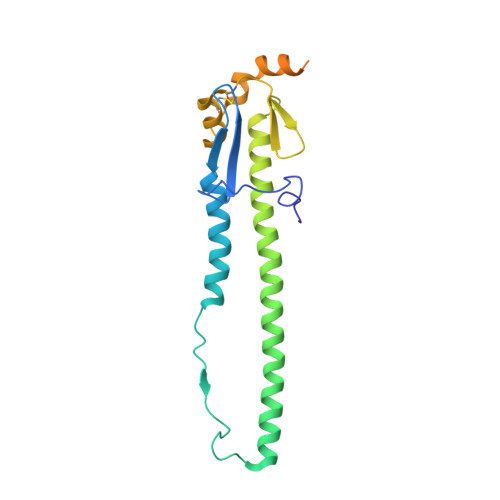
Structural and Molecular Characterization of the Hemagglutinin from the Fifth-Epidemic-Wave A(H7N9) Influenza Viruses.
Yang, H., Carney, P.J., Chang, J.C., Guo, Z., Stevens, J.(2018) J Virol 92
- PubMed: 29848588
- DOI: https://doi.org/10.1128/JVI.00375-18
- Primary Citation of Related Structures:
6D7C, 6D7U, 6D8B, 6D8D - PubMed Abstract:
The avian influenza A(H7N9) virus continues to cause human infections in China and is a major ongoing public health concern. Five epidemic waves of A(H7N9) infection have occurred since 2013, and the recent fifth epidemic wave saw the emergence of two distinct lineages with elevated numbers of human infection cases and broader geographic distribution of viral diseases compared to the first four epidemic waves. Moreover, highly pathogenic avian influenza (HPAI) A(H7N9) viruses were also isolated during the fifth epidemic wave. Here, we present a detailed structural and biochemical analysis of the surface hemagglutinin (HA) antigen from viruses isolated during this recent epidemic wave. Results highlight that, compared to the 2013 virus HAs, the fifth-wave virus HAs remained a weak binder to human glycan receptor analogs. We also studied three mutations, V177K-K184T-G219S, that were recently reported to switch a 2013 A(H7N9) HA to human-type receptor specificity. Our results indicate that these mutations could also switch the H7 HA receptor preference to a predominantly human binding specificity for both fifth-wave H7 HAs analyzed in this study. IMPORTANCE The A(H7N9) viruses circulating in China are of great public health concern. Here, we report a molecular and structural study of the major surface proteins from several recent A(H7N9) influenza viruses. Our results improve the understanding of these evolving viruses and provide important information on their receptor preference that is central to ongoing pandemic risk assessment.
Organizational Affiliation:
Influenza Division, National Center for Immunization and Respiratory Diseases, Centers for Disease Control and Prevention, Atlanta, Georgia, USA.