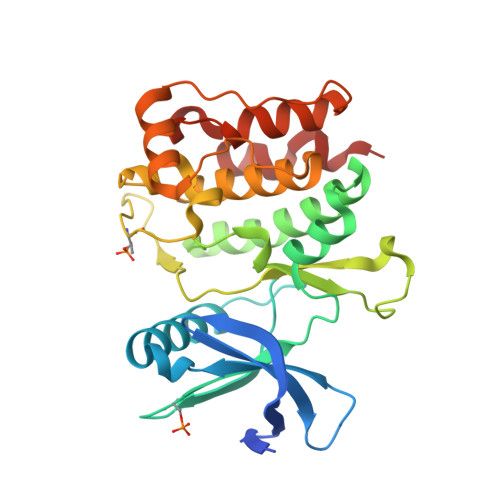
Conservation of structure, function and inhibitor binding in UNC-51-like kinase 1 and 2 (ULK1/2).
Chaikuad, A., Koschade, S.E., Stolz, A., Zivkovic, K., Pohl, C., Shaid, S., Ren, H., Lambert, L.J., Cosford, N.D.P., Brandts, C.H., Knapp, S.(2019) Biochem J 476: 875-887
- PubMed: 30782972
- DOI: https://doi.org/10.1042/BCJ20190038
- Primary Citation of Related Structures:
6QAS, 6QAT, 6QAU, 6QAV - PubMed Abstract:
Autophagy is essential for cellular homeostasis and when deregulated this survival mechanism has been associated with disease development. Inhibition of autophagy initiation by inhibiting the kinase ULK1 (Unc-51-like autophagy activating kinase 1) has been proposed as a potential cancer therapy. While inhibitors and crystal structures of ULK1 have been reported, little is known about the other closely related kinase ULK2 (Unc-51-like autophagy activating kinase 2). Here, we present the crystal structure of ULK2 in complex with ATP competitive inhibitors. Surprisingly, the ULK2 structure revealed a dimeric assembly reminiscent of dimeric arrangements of auto-activating kinases suggesting a role for this association in ULK activation. Screening of a kinase focused library of pre-clinical and clinical compounds revealed several potent ULK1/2 inhibitors and good correlation of inhibitor-binding behavior with both ULK kinases. Aurora A was identified as a major off-target of currently used ULK1 inhibitors. Autophagic flux assays demonstrated that this off-target activity by strongly inducing autophagy in different cellular systems conferred an additional layer of complexity in the interpretation of cellular data. The data presented here provide structural models and chemical starting points for the development of ULK1/2 dual inhibitors with improved selectivity for future exploitation of autophagy inhibition.
Organizational Affiliation:
Institute of Pharmaceutical Chemistry, Goethe-University Frankfurt, 60438 Frankfurt, Germany knapp@pharmchem.uni-frankfurt.de chaikuad@pharmchem.uni-frankfurt.de.