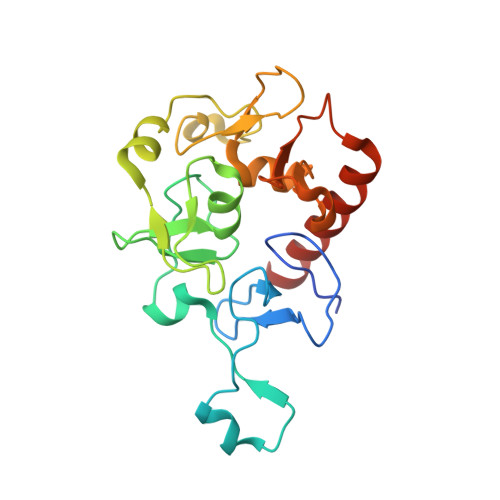
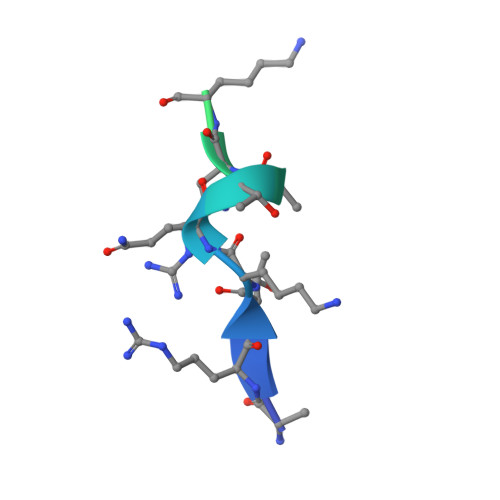
Molecular basis for bipartite recognition of histone H3 by the PZP domain of PHF14.
Zheng, S., Bi, Y., Chen, H., Gong, B., Jia, S., Li, H.(2021) Nucleic Acids Res 49: 8961-8973
- PubMed: 34365506
- DOI: https://doi.org/10.1093/nar/gkab670
- Primary Citation of Related Structures:
7D86, 7D87, 7D8A - PubMed Abstract:
Histone recognition constitutes a key epigenetic mechanism in gene regulation and cell fate decision. PHF14 is a conserved multi-PHD finger protein that has been implicated in organ development, tissue homeostasis, and tumorigenesis. Here we show that PHF14 reads unmodified histone H3(1-34) through an integrated PHD1-ZnK-PHD2 cassette (PHF14PZP). Our binding, structural and HDX-MS analyses revealed a feature of bipartite recognition, in which PHF14PZP utilizes two distinct surfaces for concurrent yet separable engagement of segments H3-Nter (e.g. 1-15) and H3-middle (e.g. 14-34) of H3(1-34). Structural studies revealed a novel histone H3 binding mode by PHD1 of PHF14PZP, in which a PHF14-unique insertion loop but not the core β-strands of a PHD finger dominates H3K4 readout. Binding studies showed that H3-PHF14PZP engagement is sensitive to modifications occurring to H3 R2, T3, K4, R8 and K23 but not K9 and K27, suggesting multiple layers of modification switch. Collectively, our work calls attention to PHF14 as a 'ground' state (unmodified) H3(1-34) reader that can be negatively regulated by active marks, thus providing molecular insights into a repressive function of PHF14 and its derepression.
Organizational Affiliation:
MOE Key Laboratory of Protein Sciences, Beijing Frontier Research Center for Biological Structure, Beijing Advanced Innovation Center for Structural Biology, Department of Basic Medical Sciences, School of Medicine, Tsinghua University, Beijing 100084, China.