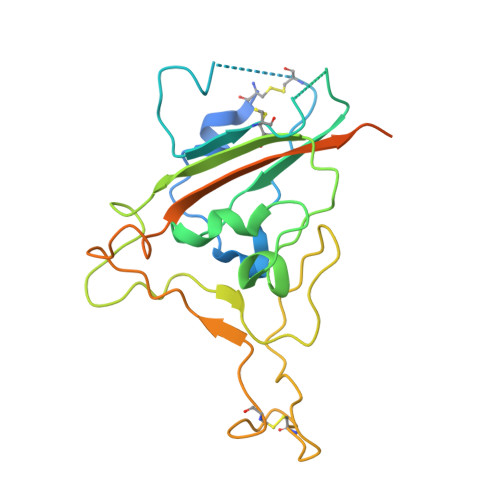
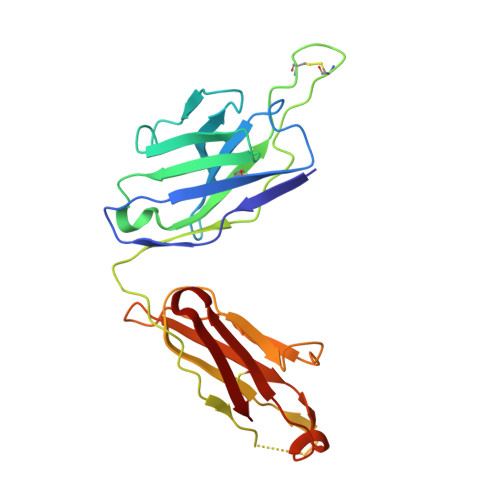
Modular basis for potent SARS-CoV-2 neutralization by a prevalent VH1-2-derived antibody class.
Rapp, M., Guo, Y., Reddem, E.R., Yu, J., Liu, L., Wang, P., Cerutti, G., Katsamba, P., Bimela, J.S., Bahna, F.A., Mannepalli, S.M., Zhang, B., Kwong, P.D., Huang, Y., Ho, D.D., Shapiro, L., Sheng, Z.(2021) Cell Rep 35: 108950-108950
- PubMed: 33794145
- DOI: https://doi.org/10.1016/j.celrep.2021.108950
- Primary Citation of Related Structures:
7L56, 7L57, 7L58, 7L5B - PubMed Abstract:
Antibodies with heavy chains that derive from the VH1-2 gene constitute some of the most potent severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)-neutralizing antibodies yet identified. To provide insight into whether these genetic similarities inform common modes of recognition, we determine the structures of the SARS-CoV-2 spike in complex with three VH1-2-derived antibodies: 2-15, 2-43, and H4. All three use VH1-2-encoded motifs to recognize the receptor-binding domain (RBD), with heavy-chain N53I-enhancing binding and light-chain tyrosines recognizing F486 RBD . Despite these similarities, class members bind both RBD-up and -down conformations of the spike, with a subset of antibodies using elongated CDRH3s to recognize glycan N343 on a neighboring RBD-a quaternary interaction accommodated by an increase in RBD separation of up to 12 Å. The VH1-2 antibody class, thus, uses modular recognition encoded by modular genetic elements to effect potent neutralization, with the VH-gene component specifying recognition of RBD and the CDRH3 component specifying quaternary interactions.
Organizational Affiliation:
Department of Biochemistry and Molecular Biophysics, Columbia University, New York, NY 10032, USA; Zuckerman Mind Brain Behavior Institute, Columbia University, New York, NY 10027, USA.