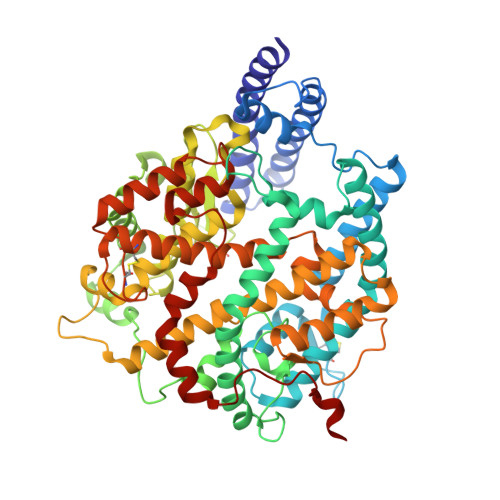
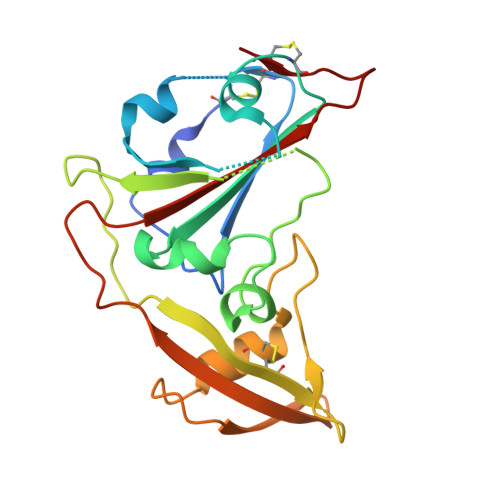
Close relatives of MERS-CoV in bats use ACE2 as their functional receptors.
Xiong, Q., Cao, L., Ma, C., Tortorici, M.A., Liu, C., Si, J., Liu, P., Gu, M., Walls, A.C., Wang, C., Shi, L., Tong, F., Huang, M., Li, J., Zhao, C., Shen, C., Chen, Y., Zhao, H., Lan, K., Corti, D., Veesler, D., Wang, X., Yan, H.(2022) Nature 612: 748-757
- PubMed: 36477529
- DOI: https://doi.org/10.1038/s41586-022-05513-3
- Primary Citation of Related Structures:
7U6R, 7WPO, 7WPZ - PubMed Abstract:
Middle East respiratory syndrome coronavirus (MERS-CoV) and several bat coronaviruses use dipeptidyl peptidase-4 (DPP4) as an entry receptor 1-4 . However, the receptor for NeoCoV-the closest known MERS-CoV relative found in bats-remains unclear 5 . Here, using a pseudotype virus entry assay, we found that NeoCoV and its close relative, PDF-2180, can efficiently bind to and use specific bat angiotensin-converting enzyme 2 (ACE2) orthologues and, less favourably, human ACE2 as entry receptors through their receptor-binding domains (RBDs) on the spike (S) proteins. Cryo-electron microscopy analysis revealed an RBD-ACE2 binding interface involving protein-glycan interactions, distinct from those of other known ACE2-using coronaviruses. We identified residues 337-342 of human ACE2 as a molecular determinant restricting NeoCoV entry, whereas a NeoCoV S pseudotyped virus containing a T510F RBD mutation efficiently entered cells expressing human ACE2. Although polyclonal SARS-CoV-2 antibodies or MERS-CoV RBD-specific nanobodies did not cross-neutralize NeoCoV or PDF-2180, an ACE2-specific antibody and two broadly neutralizing betacoronavirus antibodies efficiently inhibited these two pseudotyped viruses. We describe MERS-CoV-related viruses that use ACE2 as an entry receptor, underscoring a promiscuity of receptor use and a potential zoonotic threat.
Organizational Affiliation:
State Key Laboratory of Virology, Institute for Vaccine Research and Modern Virology Research Center, College of Life Sciences, TaiKang Center for Life and Medical Sciences, Wuhan University, Wuhan, China.