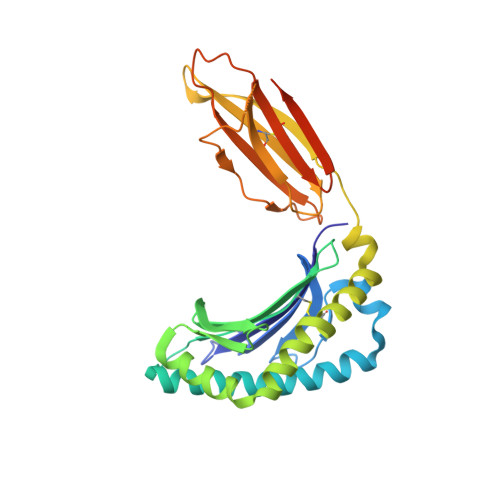
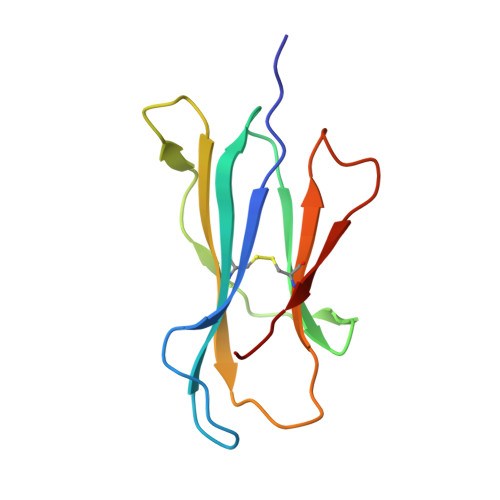
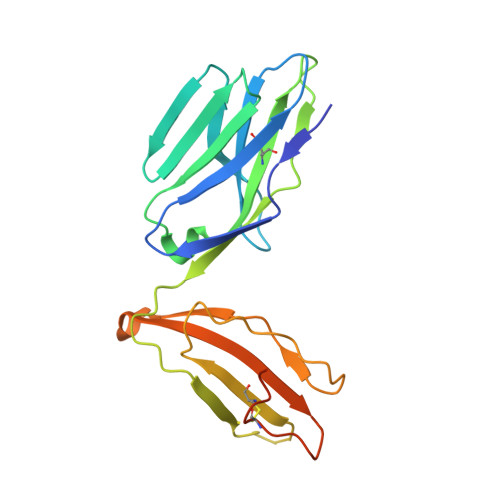
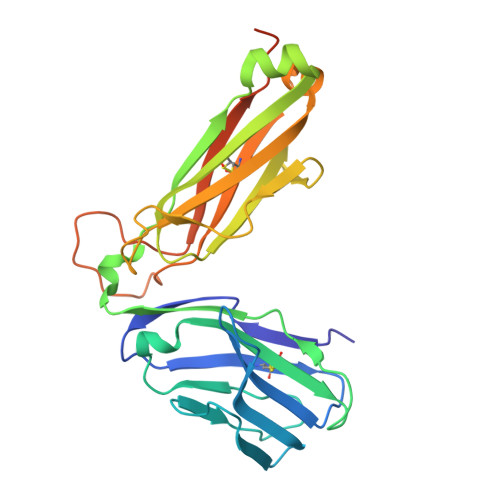
Promiscuous recognition of MR1 drives self-reactive mucosal-associated invariant T cell responses.
Chancellor, A., Alan Simmons, R., Khanolkar, R.C., Nosi, V., Beshirova, A., Berloffa, G., Colombo, R., Karuppiah, V., Pentier, J.M., Tubb, V., Ghadbane, H., Suckling, R.J., Page, K., Crean, R.M., Vacchini, A., De Gregorio, C., Schaefer, V., Constantin, D., Gligoris, T., Lloyd, A., Hock, M., Srikannathasan, V., Robinson, R.A., Besra, G.S., van der Kamp, M.W., Mori, L., Calogero, R., Cole, D.K., De Libero, G., Lepore, M.(2023) J Exp Med 220
- PubMed: 37382893
- DOI: https://doi.org/10.1084/jem.20221939
- Primary Citation of Related Structures:
7ZT2, 7ZT3, 7ZT4, 7ZT5, 7ZT7, 7ZT8, 7ZT9 - PubMed Abstract:
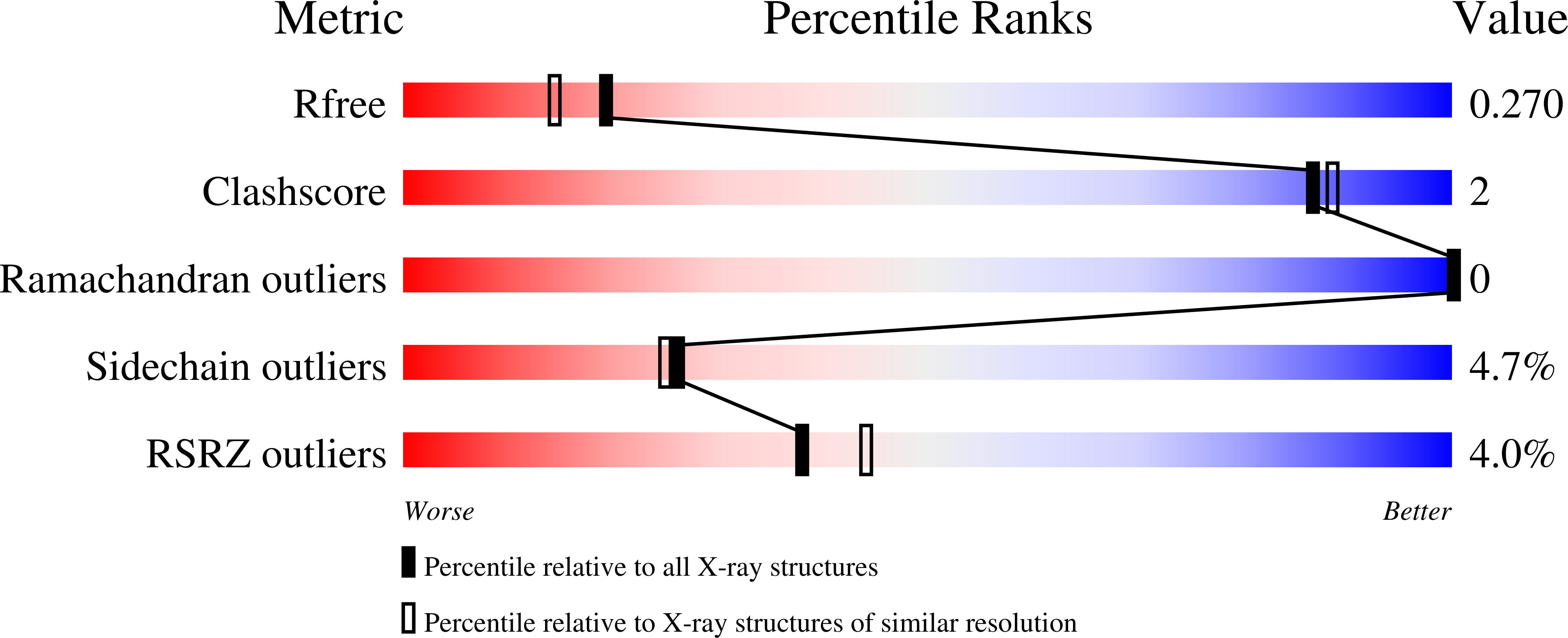
Mucosal-associated invariant T (MAIT) cells use canonical semi-invariant T cell receptors (TCR) to recognize microbial riboflavin precursors displayed by the antigen-presenting molecule MR1. The extent of MAIT TCR crossreactivity toward physiological, microbially unrelated antigens remains underexplored. We describe MAIT TCRs endowed with MR1-dependent reactivity to tumor and healthy cells in the absence of microbial metabolites. MAIT cells bearing TCRs crossreactive toward self are rare but commonly found within healthy donors and display T-helper-like functions in vitro. Experiments with MR1-tetramers loaded with distinct ligands revealed significant crossreactivity among MAIT TCRs both ex vivo and upon in vitro expansion. A canonical MAIT TCR was selected on the basis of extremely promiscuous MR1 recognition. Structural and molecular dynamic analyses associated promiscuity to unique TCRβ-chain features that were enriched within self-reactive MAIT cells of healthy individuals. Thus, self-reactive recognition of MR1 represents a functionally relevant indication of MAIT TCR crossreactivity, suggesting a potentially broader role of MAIT cells in immune homeostasis and diseases, beyond microbial immunosurveillance.
Organizational Affiliation:
Experimental Immunology, Department of Biomedicine, University Hospital Basel, University of Basel, Basel, Switzerland.