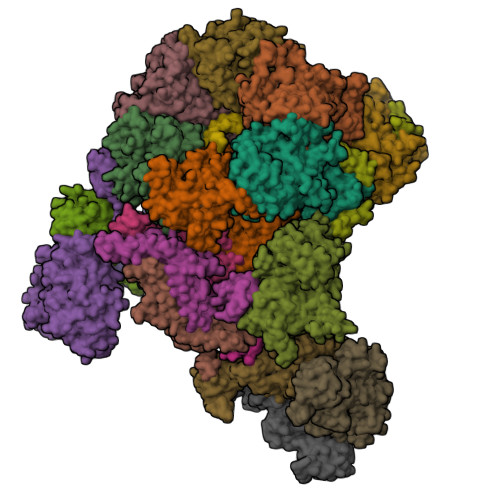
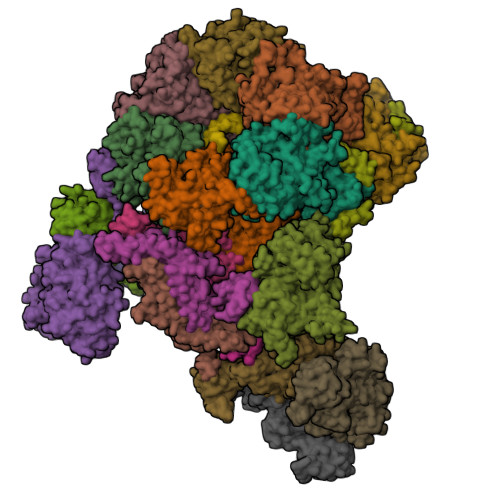
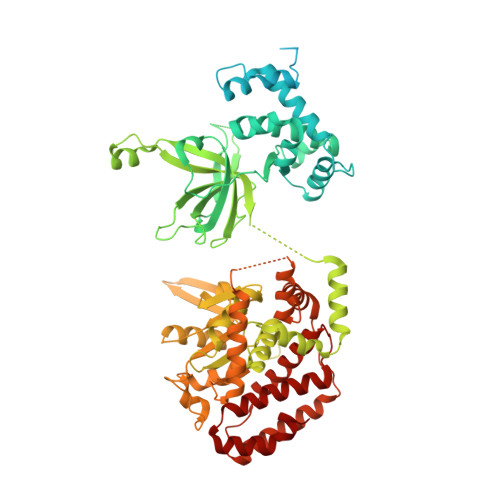
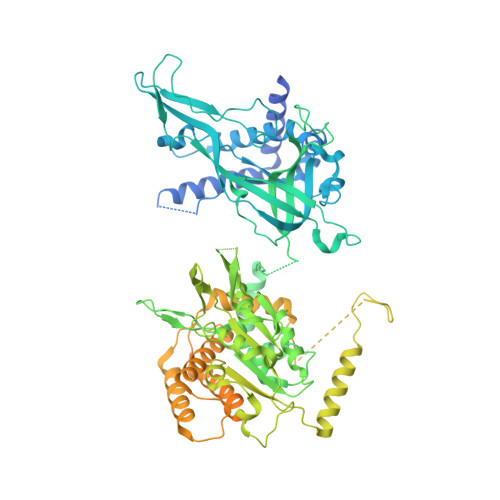
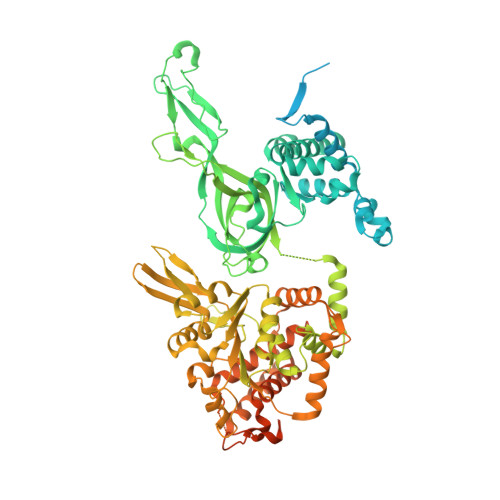
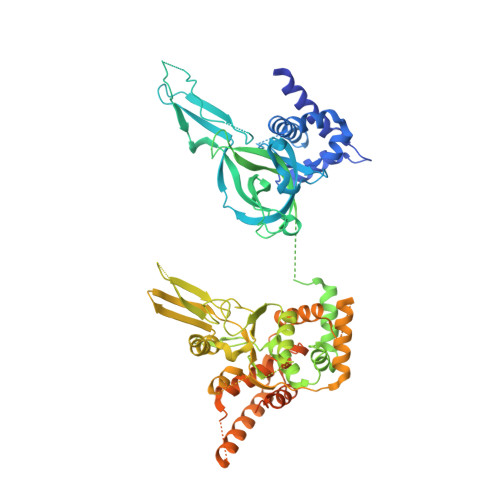
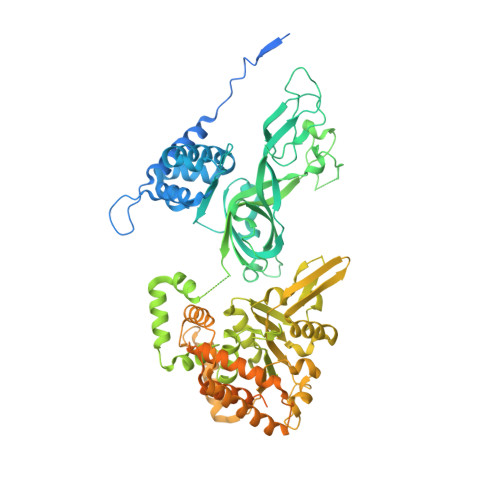
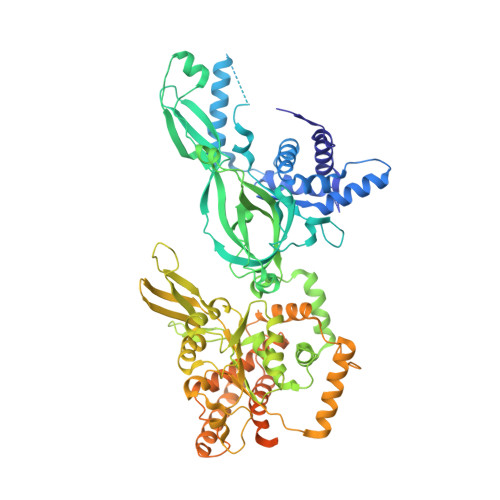
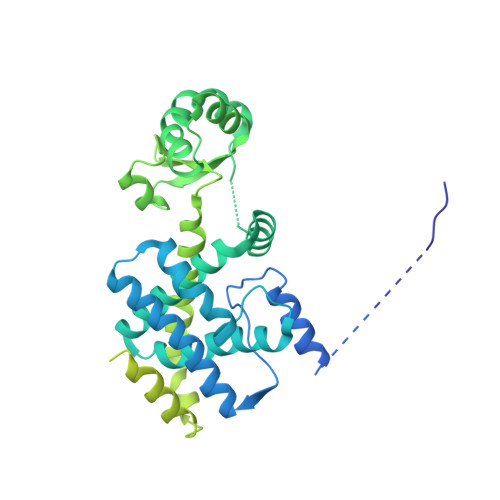
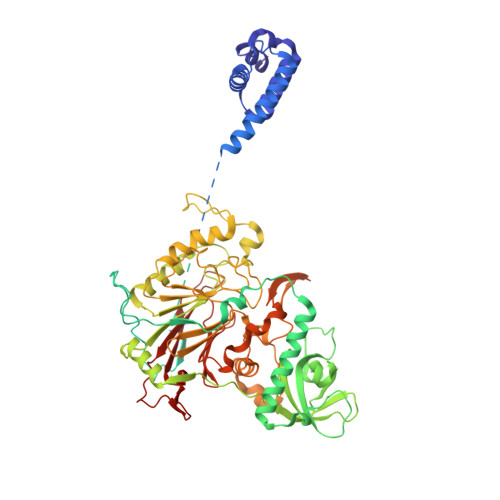
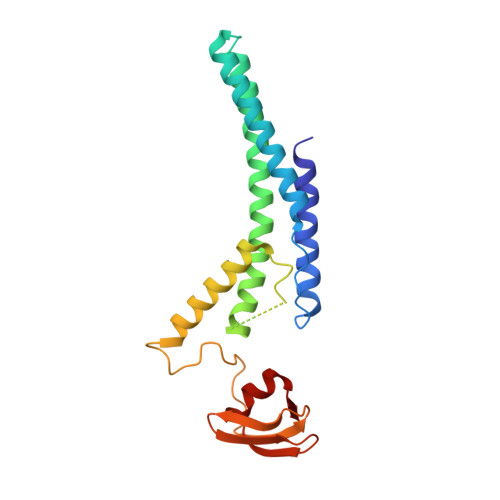
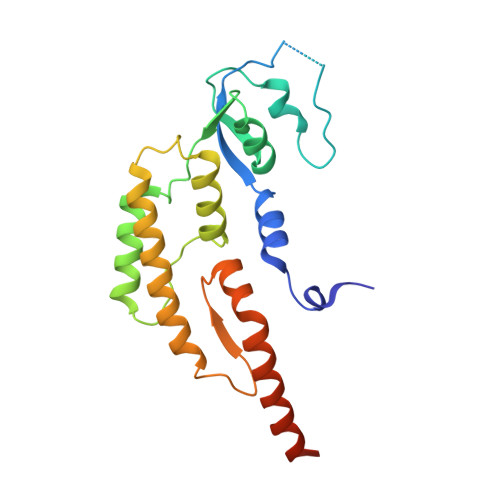
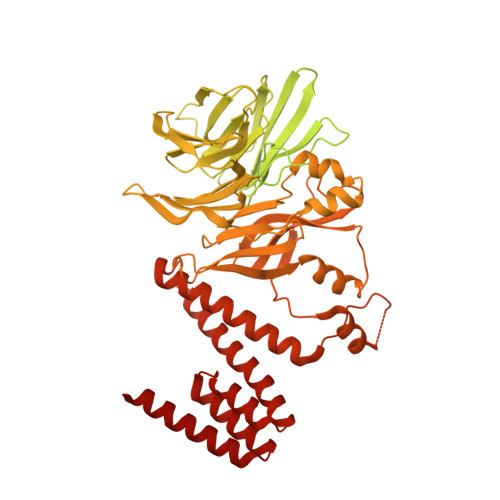
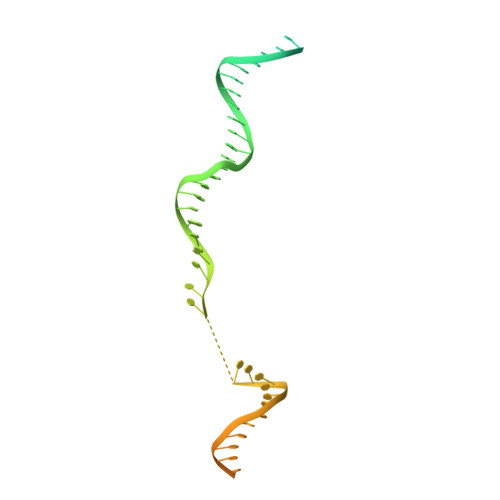
How Pol alpha-primase is targeted to replisomes to prime eukaryotic DNA replication.
Jones, M.L., Aria, V., Baris, Y., Yeeles, J.T.P.(2023) Mol Cell 83: 2911
- PubMed: 37506699
- DOI: https://doi.org/10.1016/j.molcel.2023.06.035
- Primary Citation of Related Structures:
8B9A, 8B9B, 8B9C, 8B9D - PubMed Abstract:
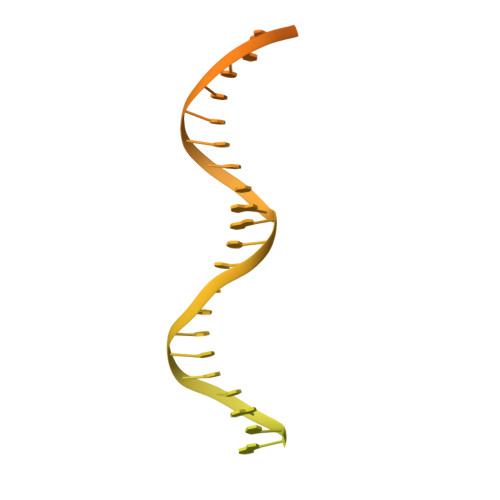
During eukaryotic DNA replication, Pol α-primase generates primers at replication origins to start leading-strand synthesis and every few hundred nucleotides during discontinuous lagging-strand replication. How Pol α-primase is targeted to replication forks to prime DNA synthesis is not fully understood. Here, by determining cryoelectron microscopy (cryo-EM) structures of budding yeast and human replisomes containing Pol α-primase, we reveal a conserved mechanism for the coordination of priming by the replisome. Pol α-primase binds directly to the leading edge of the CMG (CDC45-MCM-GINS) replicative helicase via a complex interaction network. The non-catalytic PRIM2/Pri2 subunit forms two interfaces with CMG that are critical for in vitro DNA replication and yeast cell growth. These interactions position the primase catalytic subunit PRIM1/Pri1 directly above the exit channel for lagging-strand template single-stranded DNA (ssDNA), revealing why priming occurs efficiently only on the lagging-strand template and elucidating a mechanism for Pol α-primase to overcome competition from RPA to initiate primer synthesis.
Organizational Affiliation:
MRC Laboratory of Molecular Biology, Cambridge CB2 0QH, UK.