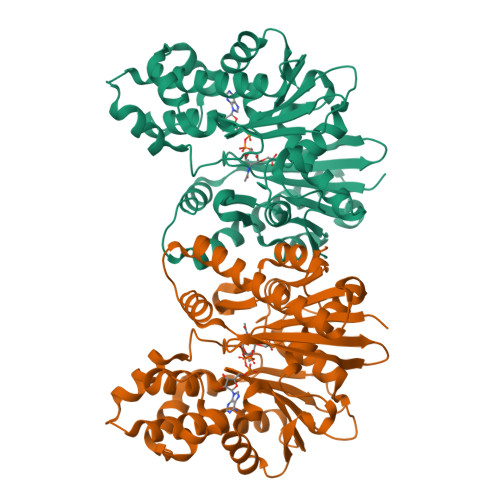
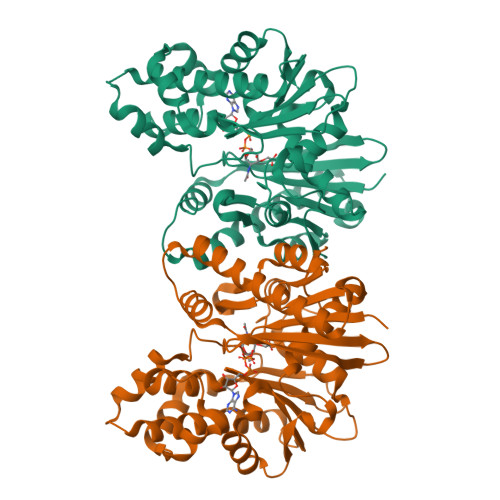
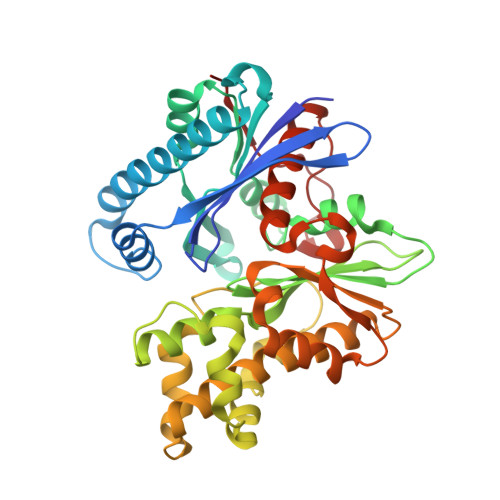
Catalytic process of anhydro-N-acetylmuramic acid kinase from Pseudomonas aeruginosa.
El-Araby, A.M., Jimenez-Faraco, E., Feltzer, R., Martin-Garcia, J.M., Karri, B.R., Ramachandran, B., Kim, C., Fisher, J.F., Hermoso, J.A., Mobashery, S.(2023) J Biological Chem 299: 105198-105198
- PubMed: 37660917
- DOI: https://doi.org/10.1016/j.jbc.2023.105198
- Primary Citation of Related Structures:
8BRE, 8C0U, 8CP9, 8CPB - PubMed Abstract:
The bacterial cell envelope is the structure with which the bacterium engages with, and is protected from, its environment. Within this envelop is a conserved peptidoglycan polymer which confers shape and strength to the cell envelop. The enzymatic processes that build, remodel, and recycle the chemical components of this cross-linked polymer are preeminent targets of antibiotics and exploratory targets for emerging antibiotic structures. We report a comprehensive kinetic and structural analysis for one such enzyme, the Pseudomonas aeruginosa anhydro-N-acetylmuramic acid (anhNAM) kinase (AnmK). AnmK is an enzyme in the peptidoglycan-recycling pathway of this pathogen. It catalyzes the pairing of hydrolytic ring opening of anhNAM with concomitant ATP-dependent phosphoryl transfer. AnmK follows a random-sequential kinetic mechanism with respect to its anhNAM and ATP substrates. Crystallographic analyses of four distinct structures (apo AnmK, AnmK:AMPPNP, AnmK:AMPPNP:anhNAM, and AnmK:ATP:anhNAM) demonstrate that both substrates enter the active site independently in an ungated conformation of the substrate subsites, with protein loops acting as gates for anhNAM binding. Catalysis occurs within a closed conformational state for the enzyme. We observe this state crystallographically using ATP-mimetic molecules. A remarkable X-ray structure for dimeric AnmK sheds light on the precatalytic and postcatalytic ternary complexes. Computational simulations in conjunction with the high-resolution X-ray structures reveal the full catalytic cycle. We further report that a P. aeruginosa strain with disrupted anmK gene is more susceptible to the β-lactam imipenem compared to the WT strain. These observations position AnmK for understanding the nexus among peptidoglycan recycling, susceptibility to antibiotics, and bacterial virulence.
Organizational Affiliation:
Department of Chemistry and Biochemistry, University of Notre Dame, Notre Dame, Indiana, USA.