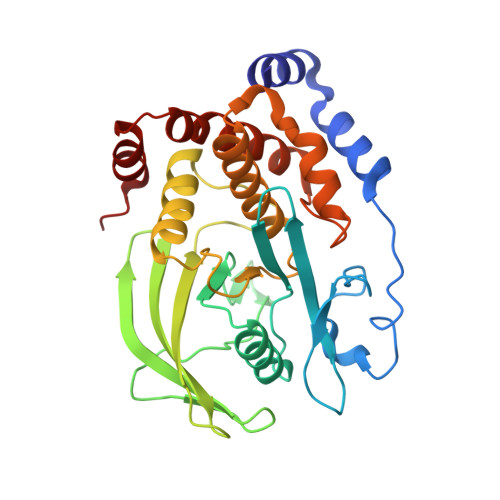
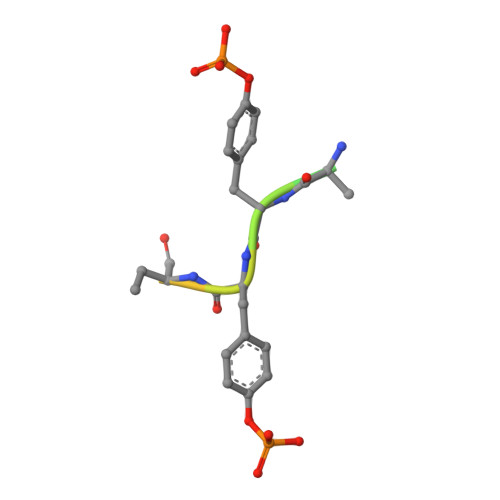
Structure guided studies of the interaction between PTP1B and JAK.
Morris, R., Keating, N., Tan, C., Chen, H., Laktyushin, A., Saiyed, T., Liau, N.P.D., Nicola, N.A., Tiganis, T., Kershaw, N.J., Babon, J.J.(2023) Commun Biol 6: 641-641
- PubMed: 37316570
- DOI: https://doi.org/10.1038/s42003-023-05020-9
- Primary Citation of Related Structures:
8EXI, 8EXJ, 8EXK, 8EXM, 8EXN, 8EYA, 8EYB, 8EYC, 8F88 - PubMed Abstract:
Protein Tyrosine Phosphatase 1B (PTP1B) is the prototypical protein tyrosine phosphatase and plays an essential role in the regulation of several kinase-driven signalling pathways. PTP1B displays a preference for bisphosphorylated substrates. Here we identify PTP1B as an inhibitor of IL-6 and show that, in vitro, it can dephosphorylate all four members of the JAK family. In order to gain a detailed understanding of the molecular mechanism of JAK dephosphorylation, we undertook a structural and biochemical analysis of the dephosphorylation reaction. We identified a product-trapping PTP1B mutant that allowed visualisation of the tyrosine and phosphate products of the reaction and a substrate-trapping mutant with a vastly decreased off-rate compared to those previously described. The latter mutant was used to determine the structure of bisphosphorylated JAK peptides bound to the enzyme active site. These structures revealed that the downstream phosphotyrosine preferentially engaged the active site, in contrast to the analogous region of IRK. Biochemical analysis confirmed this preference. In this binding mode, the previously identified second aryl binding site remains unoccupied and the non-substrate phosphotyrosine engages Arg47. Mutation of this arginine disrupts the preference for the downstream phosphotyrosine. This study reveals a previously unappreciated plasticity in how PTP1B interacts with different substrates.
Organizational Affiliation:
Walter and Eliza Hall Institute of Medical Research, 1G Royal Parade, Parkville, 3052, VIC, Australia.