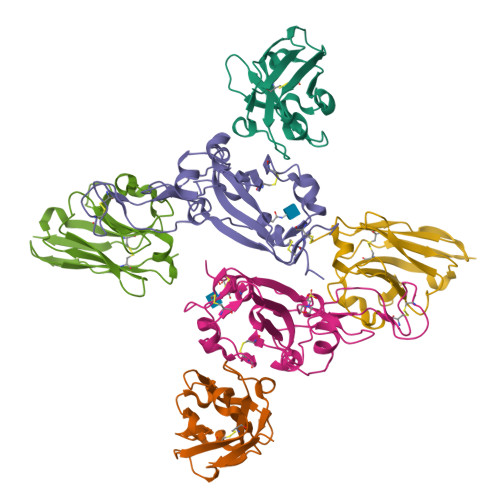
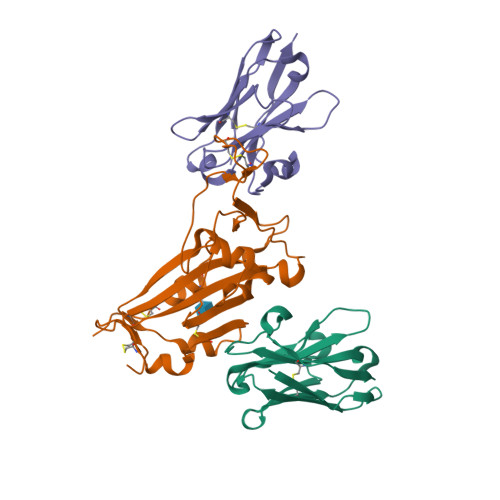
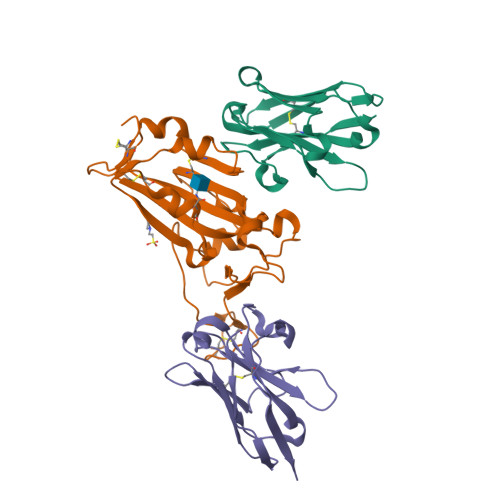
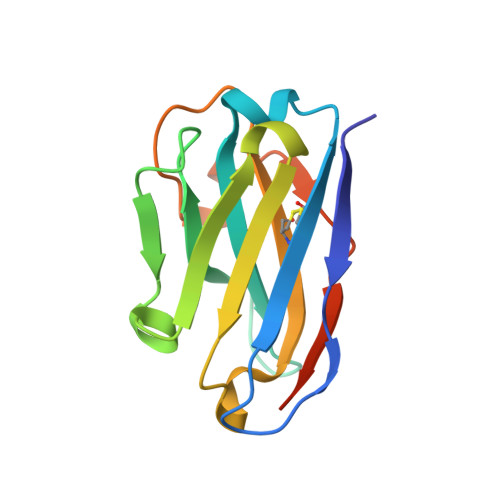
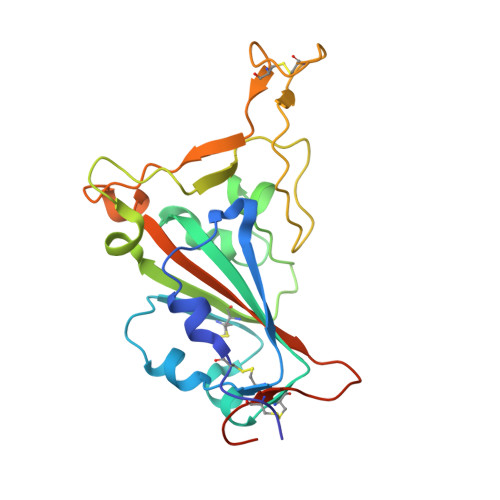
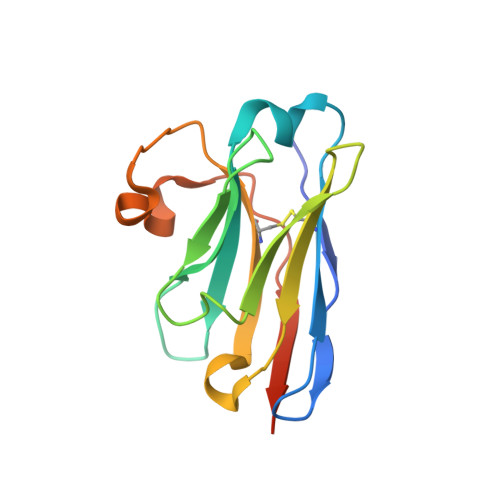
Structural and functional characterization of nanobodies that neutralize Omicron variants of SARS-CoV-2.
Cornish, K., Huo, J., Jones, L., Sharma, P., Thrush, J.W., Abdelkarim, S., Kipar, A., Ramadurai, S., Weckener, M., Mikolajek, H., Liu, S., Buckle, I., Bentley, E., Kirby, A., Han, X., Laidlaw, S.M., Hill, M., Eyssen, L., Norman, C., Le Bas, A., Clarke, J., James, W., Stewart, J.P., Carroll, M., Naismith, J.H., Owens, R.J.(2024) Open Biol 14: 230252-230252
- PubMed: 38835241
- DOI: https://doi.org/10.1098/rsob.230252
- Primary Citation of Related Structures:
8OWT, 8OWV, 8OWW, 8OYT, 8OYU - PubMed Abstract:
The Omicron strains of SARS-CoV-2 pose a significant challenge to the development of effective antibody-based treatments as immune evasion has compromised most available immune therapeutics. Therefore, in the 'arms race' with the virus, there is a continuing need to identify new biologics for the prevention or treatment of SARS-CoV-2 infections. Here, we report the isolation of nanobodies that bind to the Omicron BA.1 spike protein by screening nanobody phage display libraries previously generated from llamas immunized with either the Wuhan or Beta spike proteins. The structure and binding properties of three of these nanobodies (A8, H6 and B5-5) have been characterized in detail providing insight into their binding epitopes on the Omicron spike protein. Trimeric versions of H6 and B5-5 neutralized the SARS-CoV-2 variant of concern BA.5 both in vitro and in the hamster model of COVID-19 following nasal administration. Thus, either alone or in combination could serve as starting points for the development of new anti-viral immunotherapeutics.
Organizational Affiliation:
Structural Biology, The Rosalind Franklin Institute, Harwell Science Campus , Didcot, UK.