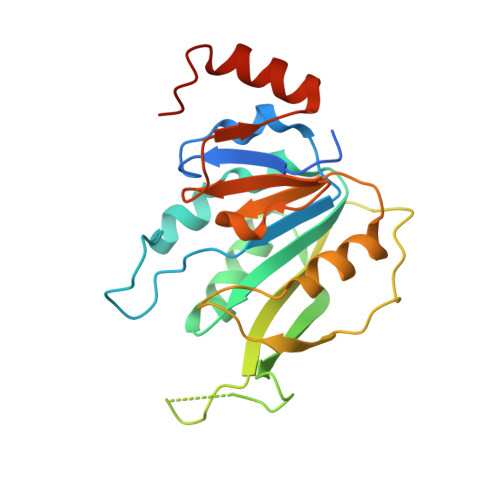
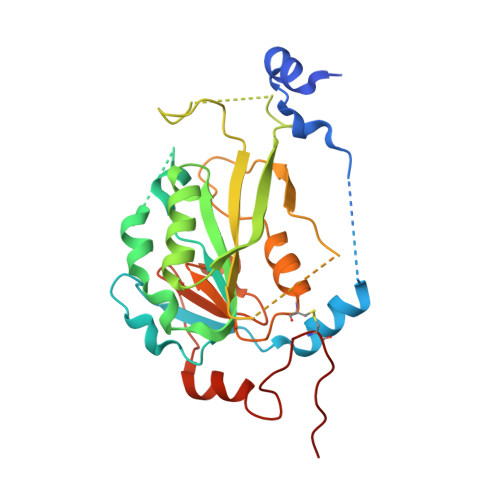
The catalytic mechanism of the RNA methyltransferase METTL3.
Corbeski, I., Vargas-Rosales, P.A., Bedi, R.K., Deng, J., Coelho, D., Braud, E., Iannazzo, L., Li, Y., Huang, D., Etheve-Quelquejeu, M., Cui, Q., Caflisch, A.(2024) Elife 12
- PubMed: 38470714
- DOI: https://doi.org/10.7554/eLife.92537
- Primary Citation of Related Structures:
8PW8, 8PW9, 8PWA, 8PWB - PubMed Abstract:
The complex of methyltransferase-like proteins 3 and 14 (METTL3-14) is the major enzyme that deposits N 6 -methyladenosine (m 6 A) modifications on messenger RNA (mRNA) in humans. METTL3-14 plays key roles in various biological processes through its methyltransferase (MTase) activity. However, little is known about its substrate recognition and methyl transfer mechanism from its cofactor and methyl donor S -adenosylmethionine (SAM). Here, we study the MTase mechanism of METTL3-14 by a combined experimental and multiscale simulation approach using bisubstrate analogues (BAs), conjugates of a SAM-like moiety connected to the N 6 -atom of adenosine. Molecular dynamics simulations based on crystal structures of METTL3-14 with BAs suggest that the Y406 side chain of METTL3 is involved in the recruitment of adenosine and release of m 6 A. A crystal structure with a BA representing the transition state of methyl transfer shows a direct involvement of the METTL3 side chains E481 and K513 in adenosine binding which is supported by mutational analysis. Quantum mechanics/molecular mechanics (QM/MM) free energy calculations indicate that methyl transfer occurs without prior deprotonation of adenosine-N 6 . Furthermore, the QM/MM calculations provide further support for the role of electrostatic contributions of E481 and K513 to catalysis. The multidisciplinary approach used here sheds light on the (co)substrate binding mechanism, catalytic step, and (co)product release, and suggests that the latter step is rate-limiting for METTL3. The atomistic information on the substrate binding and methyl transfer reaction of METTL3 can be useful for understanding the mechanisms of other RNA MTases and for the design of transition state analogues as their inhibitors.
Organizational Affiliation:
Department of Biochemistry, University of Zurich, Zurich, Switzerland.