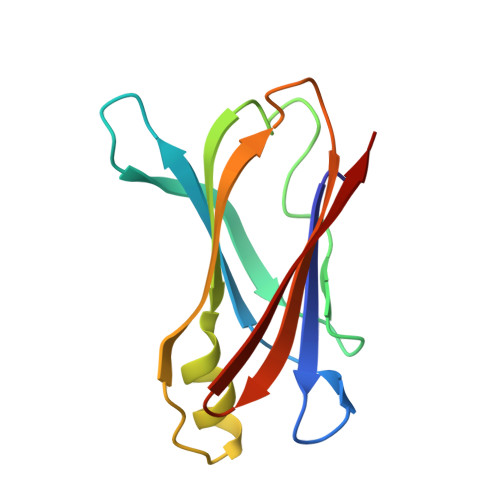
Transthyretin stability as a key factor in amyloidogenesis: X-ray analysis at atomic resolution.
Sebastiao, M.P., Lamzin, V., Saraiva, M.J., Damas, A.M.(2001) J Mol Biol 306: 733-744
- PubMed: 11243784
- DOI: https://doi.org/10.1006/jmbi.2000.4415
- Primary Citation of Related Structures:
1F86, 1FH2, 1FHN - PubMed Abstract:
Transthyretin (TTR) amyloidosis is a conformational disturbance, which, like other amyloidoses, represents a life threat. Here, we report a TTR variant, TTR Thr119Met, that has been shown to have a protective role in the development of clinical symptoms in carriers of TTR Val30Met, one of the most frequent variants among TTR amyloidosis patients. In order to understand this effect, we have determined the structures of the TTR Val30Met/Thr119Met double mutant isolated from the serum of one patient and of both the native and thyroxine complex of TTR Thr119Met. Major conclusions are: (i) new H-bonds within each monomer and monomer-monomer inter-subunit contacts, e.g. Ser117-Ser117 and Met119-Tyr114, increase protein stability, possibly leading to the protective effect of the TTR Val30Met/Thr119Met variant when compared to the single variant TTR Val30Met. (ii) The mutated residue (Met119) extends across the thyroxine binding channel inducing conformational changes that lead to closer contacts between different dimers within the tetramer. The data, at atomic resolution, were essential to detect, for the first time, the subtle changes in the inter-subunit contacts of TTR, and explain the non-amyloidogenic potential of the TTR Thr119Met variant, improving considerably current research on the TTR amyloid fibril formation pathway.
Organizational Affiliation:
Unidade de Estrutura Molecular and