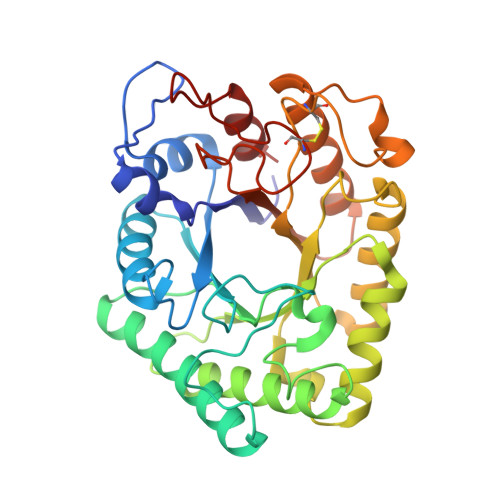
Aspergillus aculeatus beta-1,4-Galactanase: Substrate Recognition and Relations to Other Glycoside Hydrolases in Clan GH-A
Ryttersgaard, C., Leggio, L.L., Coutinho, P.M., Henrissat, B., Larsen, S.(2002) Biochemistry 41: 15135-15143
- PubMed: 12484750
- DOI: https://doi.org/10.1021/bi026238c
- Primary Citation of Related Structures:
1FHL, 1FOB - PubMed Abstract:
The three-dimensional structure of Aspergillus aculeatus beta-1,4-galactanase (AAGAL), an enzyme involved in pectin degradation, has been determined by multiple isomorphous replacement to 2.3 and 1.8 A resolution at 293 and 100 K, respectively. It represents the first known structure for a polysaccharidase with this specificity and for a member of glycoside hydrolase family 53 (GH-53). The enzyme folds into a (beta/alpha)(8) barrel with the active site cleft located at the C-terminal side of the barrel consistent with the classification of GH-53 in clan GH-A, a superfamily of enzymes with common fold and catalytic machinery but diverse specificities. Putative substrate-enzyme interactions were elucidated by modeling of beta-1,4-linked galactobioses into the possible substrate binding subsites. The structure and modeling studies identified five potential subsites for the binding of galactans, of which one is a pocket suited for accommodating the arabinan side chain in arabinogalactan, one of the natural substrates. A comparison with the substrate binding grooves of other Clan GH-A enzymes suggests that shape complementarity is crucial in determining the specificity of polysaccharidases.
- Centre for Crystallographic Studies, Department of Chemistry, University of Copenhagen, Universitetsparken 5, DK-2100 Copenhagen, Denmark.
Organizational Affiliation: