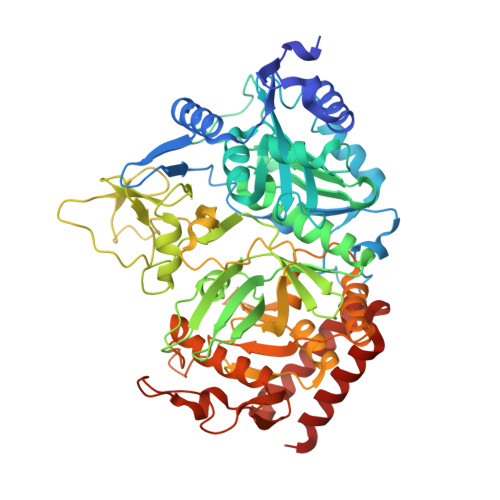
Crystal structure of human cytosolic phosphoenolpyruvate carboxykinase reveals a new GTP-binding site.
Dunten, P., Belunis, C., Crowther, R., Hollfelder, K., Kammlott, U., Levin, W., Michel, H., Ramsey, G.B., Swain, A., Weber, D., Wertheimer, S.J.(2002) J Mol Biology 316: 257-264
- PubMed: 11851336
- DOI: https://doi.org/10.1006/jmbi.2001.5364
- Primary Citation of Related Structures:
1KHB, 1KHE, 1KHF, 1KHG - PubMed Abstract:
We report crystal structures of the human enzyme phosphoenolpyruvate carboxykinase (PEPCK) with and without bound substrates. These structures are the first to be determined for a GTP-dependent PEPCK, and provide the first view of a novel GTP-binding site unique to the GTP-dependent PEPCK family. Three phenylalanine residues form the walls of the guanine-binding pocket on the enzyme's surface and, most surprisingly, one of the phenylalanine side-chains contributes to the enzyme's specificity for GTP. PEPCK catalyzes the rate-limiting step in the metabolic pathway that produces glucose from lactate and other precursors derived from the citric acid cycle. Because the gluconeogenic pathway contributes to the fasting hyperglycemia of type II diabetes, inhibitors of PEPCK may be useful in the treatment of diabetes.
- Roche Research Center, Hoffmann-La Roche Inc., Nutley, NJ 07110, USA. pete.dunten@roche.com
Organizational Affiliation: