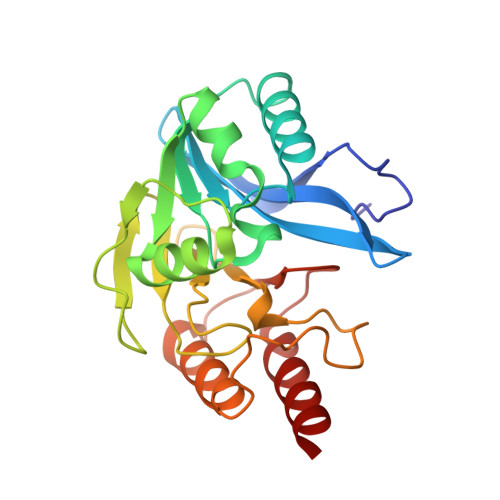
The three-dimensional structure of VIM-2, a Zn-beta-lactamase from Pseudomonas aeruginosa in its reduced and oxidised form
Garcia-Saez, I., Docquier, J.-D., Rossolini, G.M., Dideberg, O.(2008) J Mol Biology 375: 604-611
- PubMed: 18061205
- DOI: https://doi.org/10.1016/j.jmb.2007.11.012
- Primary Citation of Related Structures:
1KO2, 1KO3 - PubMed Abstract:
The crystal structures of the universally widespread metallo-beta-lactamase (MBL) Verona integron-encoded MBL (VIM)-2 from Pseudomonas aeruginosa have been solved in their native form as well as in an unexpected oxidised form. This carbapenem-hydrolysing enzyme belongs to the so-called B1 subfamily of MBLs and shares the folding of alpha beta/beta alpha sandwich, consisting of a core of beta-sheet surrounded by alpha-helices. Surprisingly, it showed a high tendency to be strongly oxidised at the catalytic cysteine located in the Cys site, Cys221, which, in the oxidised structure, becomes a cysteinesulfonic residue. Its native structure was obtained only in the presence of Tris(2-carboxyethyl)phosphine. This oxidation might be a consequence of a lower affinity for the second Zn located in the Cys site that would also explain the observed susceptibility of VIM-2 to chelating agents. This modification, if present in nature, might play a role in catalytic down-regulation. Comparison between native and oxidised VIM-2 and a predicted model of VIM-1 (which shows one residue different in the Cys site compared with VIM-2) is performed to explain the different activities and antibiotic specificities.
- Laboratoire de Cristallographie Macromoléculaire, Institut de Biologie Structurale Jean-Pierre Ebel, CNRS-Commissariat à l'Energie Atomique (CEA)-Université Joseph Fourier, 41 rue Jules Horowitz, F-38027 Grenoble Cedex 1, France. isabel.garcia@ibs.fr
Organizational Affiliation: