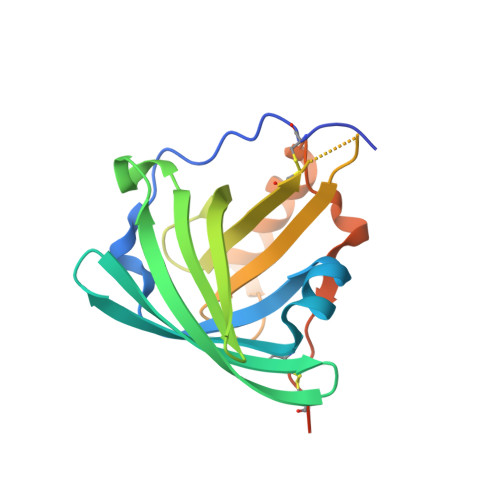
Structural mechanism of specific ligand recognition by a lipocalin tailored for the complexation of digoxigenin.
Korndoerfer, I.P., Schlehuber, S., Skerra, A.(2003) J Mol Biology 330: 385-396
- PubMed: 12823976
- DOI: https://doi.org/10.1016/s0022-2836(03)00573-4
- Primary Citation of Related Structures:
1KXO, 1LKE, 1LNM - PubMed Abstract:
DigA16 is an artificial digoxigenin-binding protein, which was derived from the bilin-binding protein, a lipocalin of Pieris brassicae, via reshaping of its natural ligand pocket. Here we report the crystal structures of DigA16 in the presence of either digoxigenin or digitoxigenin and for the apo-protein at resolutions below 1.9A. As a consequence of the altogether 17 amino acid substitutions within the binding site significant structural changes have occurred in the four loops that form the entrance to the ligand pocket on top of the structurally conserved beta-barrel framework. For example, one loop adopts a new alpha-helical backbone structure, which seems to be induced by few critical side-chain contacts. Digoxigenin becomes almost fully buried (by 95%) upon complexation, whereby specificity for the hydrophilic steroid is maintained through hydrogen-bonding networks and shape complementarity. The differential binding of the related steroid digitoxigenin is mainly governed by an internal histidine residue, whose side-chain undergoes significant induced fit. Among those amino acids that line the ligand pocket two tyrosine and one tryptophan residue provide the largest contacts. Interestingly, corresponding three side-chains are found with the same mutual orientation in the anti-digoxigenin antibody 26-10, even though the hapten orientation is quite different there and only 66% of the steroid surface is buried in the combining site. Hence, in the case of the engineered lipocalin DigA16 an example of convergent in vitro evolution is observed. Generally, the remarkable structural plasticity of the loop region and the role of polar residues in the binding site illustrate the potential of the lipocalin scaffold for the generation of specific receptor proteins towards a variety of ligands.
- Lehrstuhl für Biologische Chemie, Technische Universität München, An der Saatzucht 5, 85350, Freising-Weihenstephan, Germany.
Organizational Affiliation: