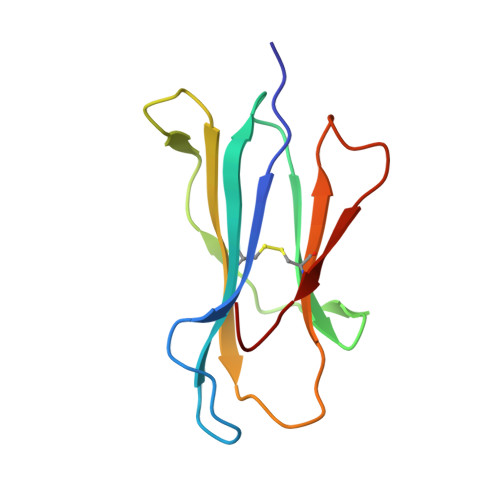
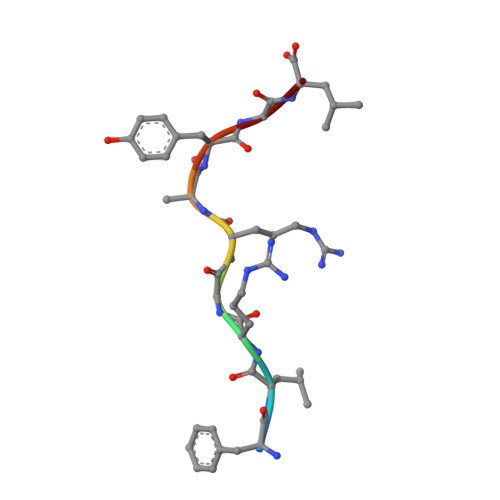
The structure of HLA-B8 complexed to an immunodominant viral determinant: peptide-induced conformational changes and a mode of MHC class I dimerization.
Kjer-Nielsen, L., Clements, C.S., Brooks, A.G., Purcell, A.W., Fontes, M.R., McCluskey, J., Rossjohn, J.(2003) J Immunol 169: 5153-5160
- PubMed: 12391232
- DOI: https://doi.org/10.4049/jimmunol.169.9.5153
- Primary Citation of Related Structures:
1M05 - PubMed Abstract:
EBV is a ubiquitous human pathogen that chronically infects up to 90% of the population. Persistent viral infection is characterized by latency and periods of viral replication that are kept in check by a strong antiviral CTL response. Despite the size of the EBV genome, CTL immunity focuses on only a few viral determinants but expands a large primary and memory response toward these epitopes. In unrelated HLA-B8(+) individuals, the response to the immunodominant latent Ag FLRGRAYGL from Epstein Barr nuclear Ag 3A is largely comprised of CTL clones with identical conserved alphabeta TCR structures. To better understand the structural correlates of Ag immunodominance and TCR selection bias, we have solved the crystal structure of the HLA-B8-FLRGRAYGL peptide complex to a resolution of 1.9 A. The structure confirms the importance of P3-Arg, P5-Arg, and P9-Leu as dominant anchor residues involved in peptide binding to HLA-B8. A bulged conformation of the bound peptide provides a structural basis for the critical role of the P7-Tyr residue in T cell recognition. The peptide also induces backbone and side-chain conformational changes in HLA-B8 that are transmitted along the peptide-binding groove in a domino effect. The HLA-B8-FLRGRAYGL complex crystallizes as a dimer in the asymmetric unit and is oriented such that both peptide ligands are projected in the same plane suggesting a higher order arrangement of MHC-peptide complexes that could be involved in formation of the class I Ag-loading complex or in T cell activation.
- Department of Microbiology and Immunology, University of Melbourne, Parkville, Victoria, Australia.
Organizational Affiliation: