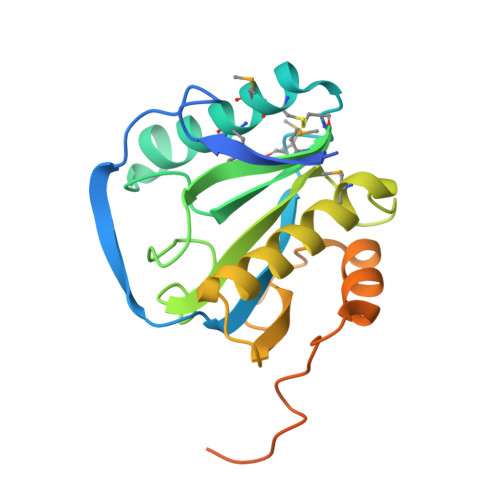
Crystal Structure of Peptidoglycan Recognition Protein Lb from Drosophila Melanogaster
Kim, M.-S., Byun, M., Oh, B.-H.(2003) Nat Immunol 4: 787
- PubMed: 12845326
- DOI: https://doi.org/10.1038/ni952
- Primary Citation of Related Structures:
1OHT - PubMed Abstract:
The family of peptidoglycan recognition proteins (PGRPs) are associated with the recognition of the peptidoglycan of microbes and subsequent activation of signaling pathways for immune response. Here the crystal structure of Drosophila PGRP-LB is determined at a resolution of 2.0 A and shows an active-site cleft with a zinc cage. Poor conservation of surface residues at the cleft predicts a widely varying individual specificity of PGRPs for molecular patterns on microbial cell walls. At the back of this cleft is a putatively conserved distinctive groove. The location and mainly hydrophobic nature of the groove indicate that the back face serves for subsequent signaling after clustering of PGRP molecules by binding to polymeric cell wall components.
- Center for Biomolecular Recognition and Division of Molecular and Life Science, Department of Life Science, Pohang University of Science and Technology, Pohang, Kyungbuk, 790-784, Korea.
Organizational Affiliation: