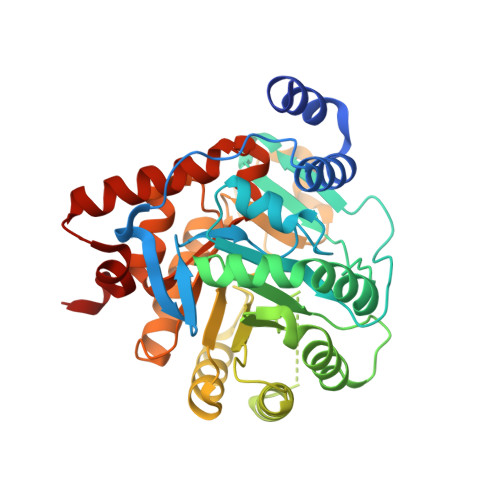
Inhibitor Binding in a Class 2 Dihydroorotate Dehydrogenase Causes Variations in the Membrane-Associated N-Terminal Domain
Hansen, M., Le Nours, J., Johansson, E., Antal, T., Ullrich, A., Loffler, M., Larsen, S.(2004) Protein Sci 13: 1031
- PubMed: 15044733
- DOI: https://doi.org/10.1110/ps.03533004
- Primary Citation of Related Structures:
1UUM, 1UUO - PubMed Abstract:
The flavin enzyme dihydroorotate dehydrogenase (DHOD; EC 1.3.99.11) catalyzes the oxidation of dihydroorotate to orotate, the fourth step in the de novo pyrimidine biosynthesis of UMP. The enzyme is a promising target for drug design in different biological and clinical applications for cancer and arthritis. The first crystal structure of the class 2 dihydroorotate dehydrogenase from rat has been determined in complex with its two inhibitors brequinar and atovaquone. These inhibitors have shown promising results as anti-proliferative, immunosuppressive, and antiparasitic agents. A unique feature of the class 2 DHODs is their N-terminal extension, which folds into a separate domain comprising two alpha-helices. This domain serves as the binding site for the two inhibitors and the respiratory quinones acting as the second substrate for the class 2 DHODs. The orientation of the first N-terminal helix is very different in the two complexes of rat DHOD (DHODR). Binding of atovaquone causes a 12 A movement of the first residue in the first alpha-helix. Based on the information from the two structures of DHODR, a model for binding of the quinone and the residues important for the interactions could be defined. His 56 and Arg 136, which are fully conserved in all class 2 DHODs, seem to play a key role in the interaction with the electron acceptor. The differences between the membrane-bound rat DHOD and membrane-associated class 2 DHODs exemplified by the Escherichia coli DHOD has been investigated by GRID computations of the hydrophobic probes predicted to interact with the membrane.
Organizational Affiliation:
Centre for Crystallographic Studies, Department of Chemistry, University of Copenhagen, Universitetsparken 5, DK-2100 Copenhagen, Denmark.