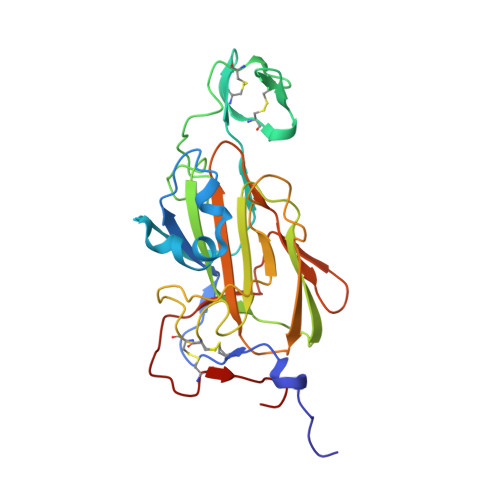
Structural Basis of Flocculin-Mediated Social Behavior in Yeast
Veelders, M., Brueckner, S., Ott, D., Unverzagt, C., Moesch, H.-U., Essen, L.-O.(2010) Proc Natl Acad Sci U S A 107: 22511
- PubMed: 21149680
- DOI: https://doi.org/10.1073/pnas.1013210108
- Primary Citation of Related Structures:
2XJP, 2XJQ, 2XJR, 2XJS, 2XJT, 2XJU, 2XJV - PubMed Abstract:
In the budding yeast Saccharomyces cerevisiae, self-recognition and the thereby promoted aggregation of thousands of cells into protective flocs is mediated by a family of cell-surface adhesins, the flocculins (Flo). Based on this social behavior FLO genes fulfill the definition of "greenbeard" genes, which direct cooperation toward other carriers of the same gene. The process of flocculation plays an eminent role in the food industry for the production of beer and wine. However, the precise mode of flocculin-mediated surface recognition and the exact structure of cognate ligands have remained elusive. Here, we present structures of the adhesion domain of a flocculin complexed to its cognate ligands derived from yeast high-mannose oligosaccharides at resolutions up to 0.95 Å. Besides a PA14-like architecture, the Flo5A domain reveals a previously undescribed lectin fold that utilizes a unique DcisD calcium-binding motif for carbohydrate binding and that is widely spread among pro- and eukaryotes. Given the high abundance of high-mannose oligosaccharides in yeast cell walls, the Flo5A structure suggests a model for recognition, where social non-self- instead of unsocial self-interactions are favored.
Organizational Affiliation:
Department of Chemistry/Biochemistry, Philipps-Universität Marburg, Hans-Meerwein-Strasse, D-35032 Marburg, Germany.