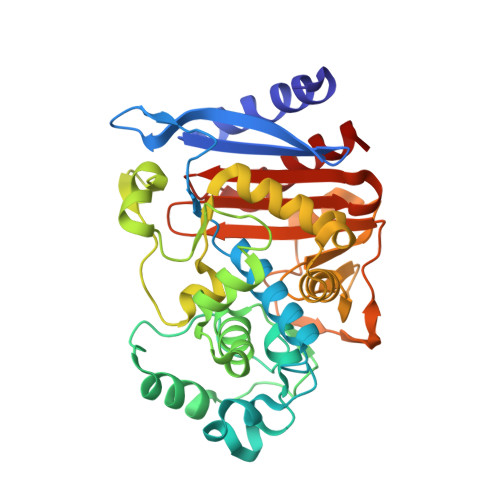
Structure of AmpC beta-lactamase (AmpCD) from an Escherichia coli clinical isolate with a tripeptide deletion (Gly286-Ser287-Asp288) in the H10 helix
Yamaguchi, Y., Sato, G., Yamagata, Y., Doi, Y., Wachino, J., Arakawa, Y., Matsuda, K., Kurosaki, H.(2009) Acta Crystallogr Sect F Struct Biol Cryst Commun 65: 540-543
- PubMed: 19478427
- DOI: https://doi.org/10.1107/S1744309109014249
- Primary Citation of Related Structures:
2ZJ9 - PubMed Abstract:
The X-ray crystal structure of AmpC beta-lactamase (AmpC(D)) with a tripeptide deletion (Gly286-Ser287-Asp288) produced by Escherichia coli HKY28, a ceftazidime-resistant strain, was determined at a resolution of 1.7 A. The structure of AmpC(D) suggests that the tripeptide deletion at positions 286-288 located in the H10 helix causes a structural change of the Asn289-Asn294 region from the alpha-helix present in the native AmpC beta-lactamase of E. coli to a loop structure, which results in a widening of the substrate-binding site.
- Environmental Safety Center, Kumamoto University, Kumamoto, Japan. yyamagu@gpo.kumamoto-u.ac.jp
Organizational Affiliation: