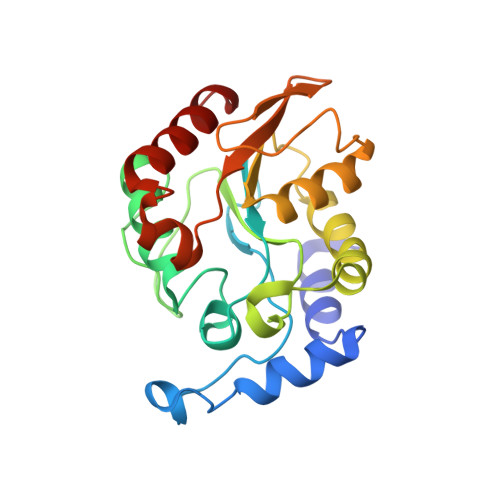
Crystal structure of family 5 uracil-DNA glycosylase bound to DNA.
Kosaka, H., Hoseki, J., Nakagawa, N., Kuramitsu, S., Masui, R.(2007) J Mol Biology 373: 839-850
- PubMed: 17870091
- DOI: https://doi.org/10.1016/j.jmb.2007.08.022
- Primary Citation of Related Structures:
2D3Y, 2DDG, 2DEM - PubMed Abstract:
Uracil-DNA glycosylase (UDG) removes uracil generated by the deamination of cytosine or misincorporation of deoxyuridine monophosphate. Within the UDG superfamily, a fifth UDG family lacks a polar residue in the active-site motif, which mediates the hydrolysis of the glycosidic bond by activation of a water molecule in UDG families 1-4. We have determined the crystal structure of a novel family 5 UDG from Thermus thermophilus HB8 complexed with DNA containing an abasic site. The active-site structure suggests this enzyme uses both steric force and water activation for its excision reaction. A conserved asparagine residue acts as a ligand to the catalytic water molecule. The structure also implies that another water molecule acts as a barrier during substrate recognition. Based on no significant open-closed conformational change upon binding to DNA, we propose a "slide-in" mechanism for initial damage recognition.
- Department of Biological Sciences, Graduate School of Science, Osaka University, 1-1 Machikaneyama-cho, Toyonaka, Osaka 560-0043, Japan.
Organizational Affiliation: