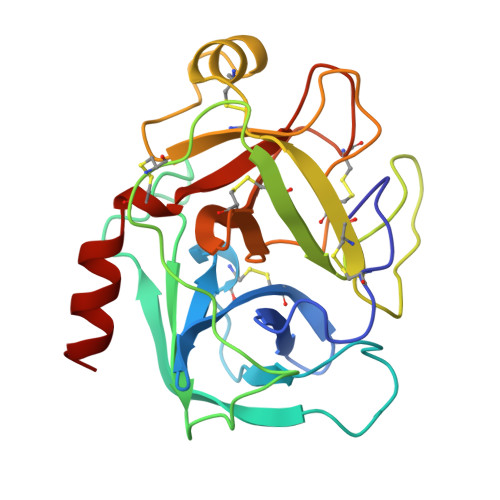
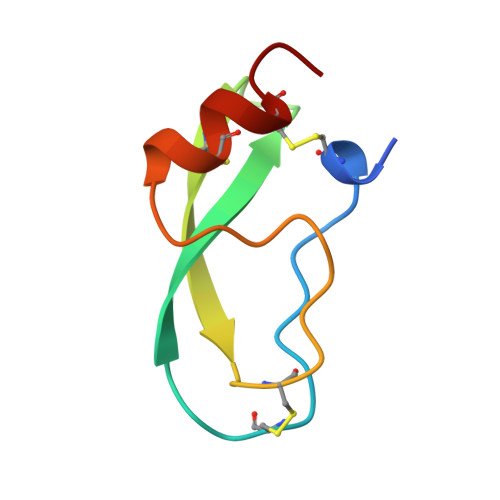
The crystal structures of the complexes between bovine beta-trypsin and ten P1 variants of BPTI.
Helland, R., Otlewski, J., Sundheim, O., Dadlez, M., Smalas, A.O.(1999) J Mol Biology 287: 923-942
- PubMed: 10222201
- DOI: https://doi.org/10.1006/jmbi.1999.2654
- Primary Citation of Related Structures:
3BTD, 3BTE, 3BTF, 3BTG, 3BTH, 3BTK, 3BTM, 3BTQ, 3BTT, 3BTW - PubMed Abstract:
The high-resolution X-ray structures have been determined for ten complexes formed between bovine beta-trypsin and P1 variants (Gly, Asp, Glu, Gln, Thr, Met, Lys, His, Phe, Trp) of bovine pancreatic trypsin inhibitor (BPTI). All the complexes were crystallised from the same conditions. The structures of the P1 variants Asp, Glu, Gln and Thr, are reported here for the first time in complex with any serine proteinase. The resolution of the structures ranged from 1.75 to 2.05 A and the R-factors were about 19-20 %. The association constants of the mutants ranged from 1.5x10(4) to 1.7x10(13) M-1. All the structures could be fitted into well-defined electron density, and all had very similar global conformations. All the P1 mutant side-chains could be accomodated at the primary binding site, but relative to the P1 Lys, there were small local changes within the P1-S1 interaction site. These comprised: (1) changes in the number and dynamics of water molecules inside the pocket; (2) multiple conformations and non-optimal dihedral angles for some of the P1 side-chains, Ser190 and Gln192; and (3) changes in temperature factors of the pocket walls as well as the introduced P1 side-chain. Binding of the cognate P1 Lys is characterised by almost optimal dihedral angles, hydrogen bonding distances and angles, in addition to considerably lower temperature factors. Thus, the trypsin S1 pocket seems to be designed particularly for lysine binding.
- Department of Chemistry, University of Tromsø, Tromsø, 9037, Norway.
Organizational Affiliation: