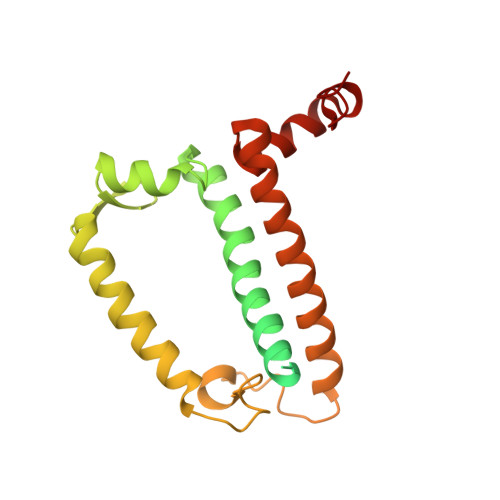
Structural insights into energy regulation of light-harvesting complex CP29 from spinach.
Pan, X., Li, M., Wan, T., Wang, L., Jia, C., Hou, Z., Zhao, X., Zhang, J., Chang, W.(2011) Nat Struct Mol Biol 18: 309-315
- PubMed: 21297637
- DOI: https://doi.org/10.1038/nsmb.2008
- Primary Citation of Related Structures:
3PL9 - PubMed Abstract:
CP29, one of the minor light-harvesting complexes of higher-plant photosystem II, absorbs and transfers solar energy for photosynthesis and also has important roles in photoprotection. We have solved the crystal structure of spinach CP29 at 2.80-Å resolution. Each CP29 monomer contains 13 chlorophyll and 3 carotenoid molecules, which differs considerably from the major light-harvesting complex LHCII and the previously proposed CP29 model. The 13 chlorophyll-binding sites are assigned as eight chlorophyll a sites, four chlorophyll b and one putative mixed site occupied by both chlorophylls a and b. Based on the present X-ray structure, an integrated pigment network in CP29 is constructed. Two special clusters of pigment molecules, namely a615-a611-a612-Lut and Vio(Zea)-a603-a609, have been identified and might function as potential energy-quenching centers and as the exit or entrance in energy-transfer pathways.
- National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing, China.
Organizational Affiliation: