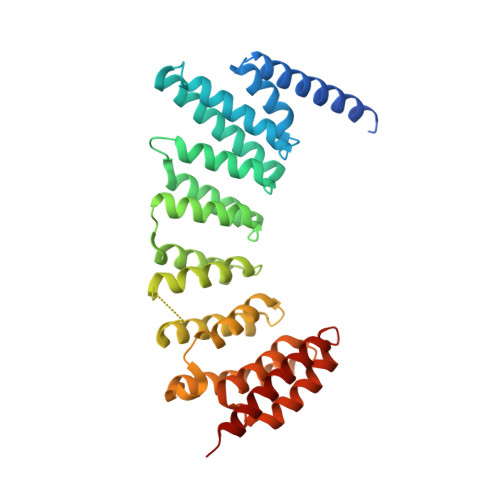
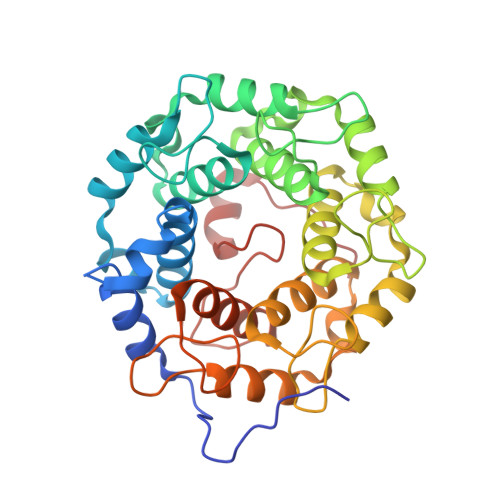
Structure-Guided Development of Selective RabGGTase Inhibitors.
Bon, R.S., Guo, Z., Stigter, E.A., Wetzel, S., Menninger, S., Wolf, A., Choidas, A., Alexandrov, K., Blankenfeldt, W., Goody, R.S., Waldmann, H.(2011) Angew Chem Int Ed Engl 50: 4957-4961