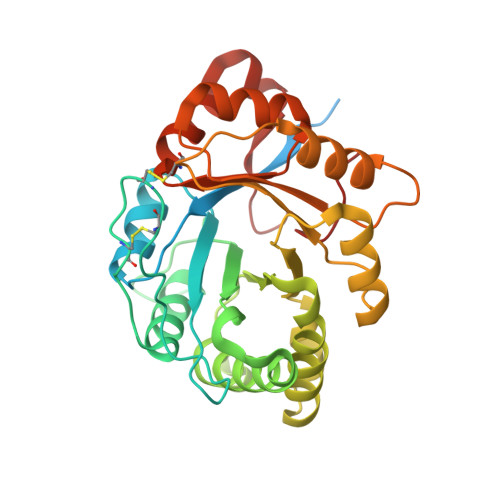
Structure of a novel class II phospholipase D: Catalytic cleft is modified by a disulphide bridge.
de Giuseppe, P.O., Ullah, A., Silva, D.T., Gremski, L.H., Wille, A.C., Chaves Moreira, D., Ribeiro, A.S., Chaim, O.M., Murakami, M.T., Veiga, S.S., Arni, R.K.(2011) Biochem Biophys Res Commun 409: 622-627
- PubMed: 21616057
- DOI: https://doi.org/10.1016/j.bbrc.2011.05.053
- Primary Citation of Related Structures:
3RLH - PubMed Abstract:
Phospholipases D (PLDs) are principally responsible for the local and systemic effects of Loxosceles envenomation including dermonecrosis and hemolysis. Despite their clinical relevance in loxoscelism, to date, only the SMase I from Loxosceles laeta, a class I member, has been structurally characterized. The crystal structure of a class II member from Loxosceles intermedia venom has been determined at 1.7Å resolution. Structural comparison to the class I member showed that the presence of an additional disulphide bridge which links the catalytic loop to the flexible loop significantly changes the volume and shape of the catalytic cleft. An examination of the crystal structures of PLD homologues in the presence of low molecular weight compounds at their active sites suggests the existence of a ligand-dependent rotamer conformation of the highly conserved residue Trp230 (equivalent to Trp192 in the glycerophosphodiester phosphodiesterase from Thermus thermophofilus, PDB code: 1VD6) indicating its role in substrate binding in both enzymes. Sequence and structural analyses suggest that the reduced sphingomyelinase activity observed in some class IIb PLDs is probably due to point mutations which lead to a different substrate preference.
- Laboratório Nacional de Biociências, Centro Nacional de Pesquisa em Energia e Materiais, Campinas, 13083-970 SP, Brazil.
Organizational Affiliation: