Human and mouse type I natural killer T cell antigen receptors exhibit different fine specificities for CD1d-antigen complex
Wun, K.S., Ross, F., Patel, O., Besra, G.S., Porcelli, S.A., Richardson, S.K., Keshipeddy, S., Howell, A.R., Godfrey, D.I., Rossjohn, J.(2012) J Biological Chem 287: 39139-39148
- PubMed: 22995911
- DOI: https://doi.org/10.1074/jbc.M112.412320
- Primary Citation of Related Structures:
3VWJ, 3VWK - PubMed Abstract:
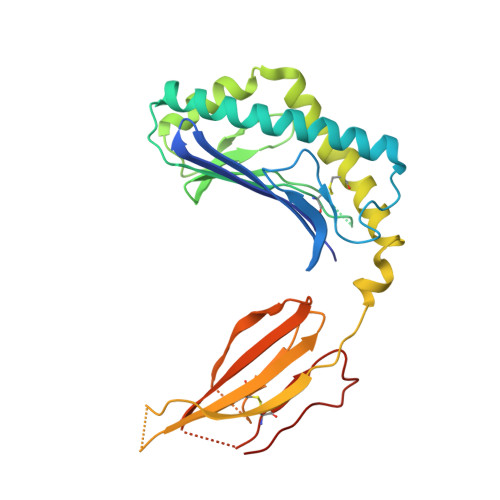
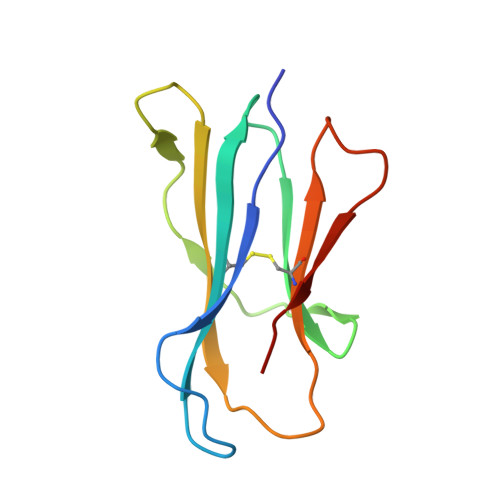
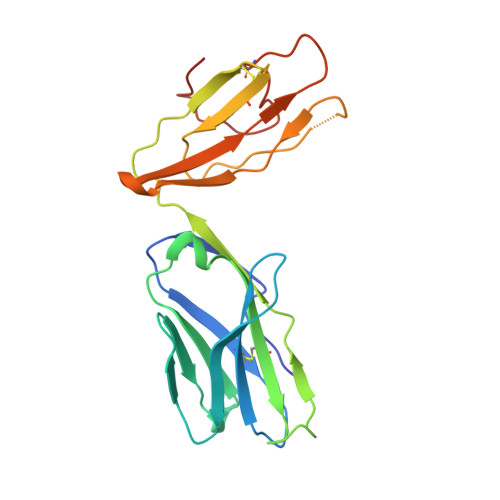
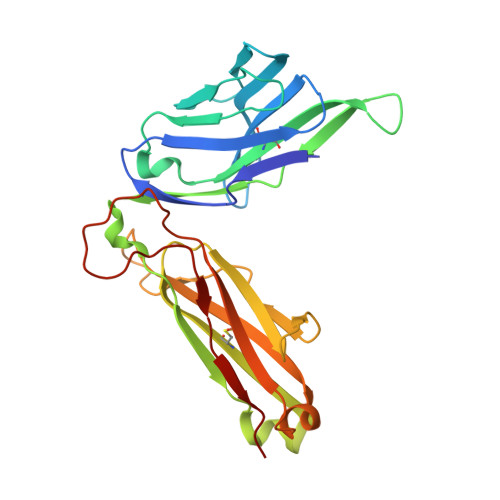
Human and mouse type I natural killer T (NKT) cells respond to a variety of CD1d-restricted glycolipid antigens (Ags), with their NKT cell antigen receptors (NKT TCRs) exhibiting reciprocal cross-species reactivity that is underpinned by a conserved NKT TCR-CD1d-Ag docking mode. Within this common docking footprint, the NKT TCR recognizes, to varying degrees of affinity, a range of Ags. Presently, it is unclear whether the human NKT TCRs will mirror the generalities underpinning the fine specificity of the mouse NKT TCR-CD1d-Ag interaction. Here, we assessed human NKT TCR recognition against altered glycolipid ligands of α-galactosylceramide (α-GalCer) and have determined the structures of a human NKT TCR in complex with CD1d-4',4″-deoxy-α-GalCer and CD1d-α-GalCer with a shorter, di-unsaturated acyl chain (C20:2). Altered glycolipid ligands with acyl chain modifications did not affect the affinity of the human NKT TCR-CD1d-Ag interaction. Surprisingly, human NKT TCR recognition is more tolerant to modifications at the 4'-OH position in comparison with the 3'-OH position of α-GalCer, which contrasts the fine specificity of the mouse NKT TCR-CD1d-Ag recognition (4'-OH > 3'-OH). The fine specificity differences between human and mouse NKT TCRs was attributable to differing interactions between the respective complementarity-determining region 1α loops and the Ag. Accordingly, germline encoded fine-specificity differences underpin human and mouse type I NKT TCR interactions, which is an important consideration for therapeutic development and NKT cell physiology.
- Australian Research Council (ARC) Centre of Excellence in Structural and Functional Microbial Genomics, Department of Biochemistry and Molecular Biology, School of Biomedical Sciences, Monash University, Clayton, Victoria 3800, Australia.
Organizational Affiliation: