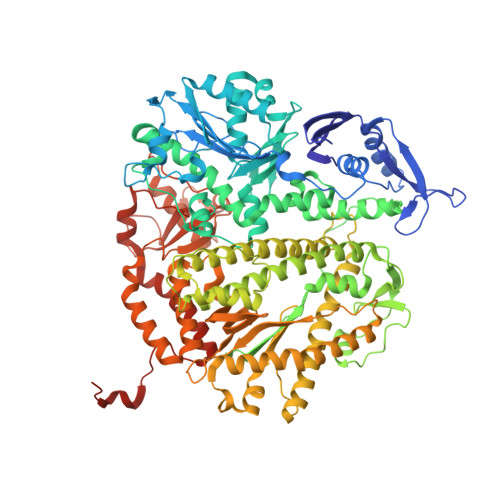
Phosphonoformic acid inhibits viral replication by trapping the closed form of the DNA polymerase.
Zahn, K.E., Tchesnokov, E.P., Gotte, M., Doublie, S.(2011) J Biol Chem 286: 25246-25255
- PubMed: 21566148
- DOI: https://doi.org/10.1074/jbc.M111.248864
- Primary Citation of Related Structures:
3KD1, 3KD5 - PubMed Abstract:
Phosphonoformic acid (PFA, foscarnet) belongs to a class of antiviral drugs that inhibit the human cytomegalovirus DNA polymerase (UL54) by mimicking the pyrophosphate leaving group of the nucleotide transfer reaction. Difficulties expressing UL54 have hampered investigation of the precise structural requirements rendering inhibition by this drug. However, a previously engineered chimeric DNA polymerase, constructed by mutating the homologous polymerase from bacteriophage RB69 (gp43) to express several variable elements from UL54, can bypass this obstacle because of its favorable expression and acquired sensitivity to PFA (Tchesnokov, E. P., Obikhod, A., Schinazi, R. F., and Götte, M. (2008) J. Biol. Chem. 283, 34218-34228). Here, we compare two crystal structures that depict the chimeric DNA polymerase with and without PFA bound. PFA is visualized for the first time in the active site of a DNA polymerase, where interactions are resolved between the PP(i) mimic and two basic residues absolutely conserved in the fingers domain of family B polymerases. PFA also chelates metal ion B, the cation that contacts the triphosphate tail of the incoming nucleotide. These DNA complexes utilize a primer-template pair enzymatically chain-terminated by incorporation of acyclo-GMP, the phosphorylated form of the anti-herpes drug acyclovir. We postulate that the V478W mutation present in the chimera is critical in that it pushes the fingers domain to more readily adopt the closed conformation whether or not the drug is bound. The closed state of the fingers domain traps the variant polymerase in the untranslocated state and increases affinity for PFA. This finding provides a model for the mechanism of UL54 stalling by PFA.
Organizational Affiliation:
Department of Microbiology and Molecular Genetics, University of Vermont, Burlington, Vermont 05405, USA.