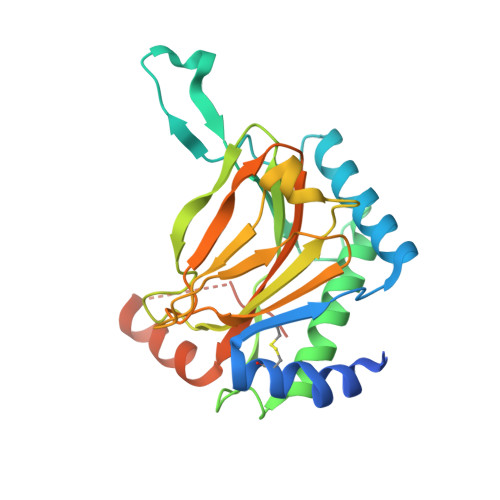
Selective small molecule probes for the hypoxia inducible factor (HIF) prolyl hydroxylases.
Chowdhury, R., Candela-Lena, J.I., Chan, M.C., Greenald, D.J., Yeoh, K.K., Tian, Y.M., McDonough, M.A., Tumber, A., Rose, N.R., Conejo-Garcia, A., Demetriades, M., Mathavan, S., Kawamura, A., Lee, M.K., van Eeden, F., Pugh, C.W., Ratcliffe, P.J., Schofield, C.J.(2013) ACS Chem Biol 8: 1488-1496
- PubMed: 23683440
- DOI: https://doi.org/10.1021/cb400088q
- Primary Citation of Related Structures:
4BQW, 4BQX, 4BQY - PubMed Abstract:
The hypoxia inducible factor (HIF) system is central to the signaling of low oxygen (hypoxia) in animals. The levels of HIF-α isoforms are regulated in an oxygen-dependent manner by the activity of the HIF prolyl-hydroxylases (PHD or EGLN enzymes), which are Fe(II) and 2-oxoglutarate (2OG) dependent oxygenases. Here, we describe biochemical, crystallographic, cellular profiling, and animal studies on PHD inhibitors including selectivity studies using a representative set of human 2OG oxygenases. We identify suitable probe compounds for use in studies on the functional effects of PHD inhibition in cells and in animals.
- Department of Chemistry, Chemistry Research Laboratory, University of Oxford , Mansfield Road, Oxford, OX1 3TA, United Kingdom.
Organizational Affiliation: