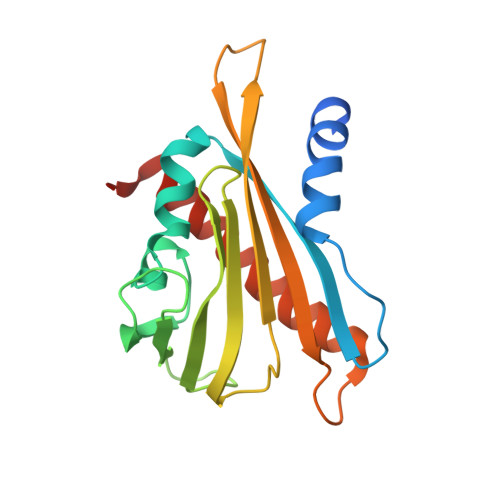
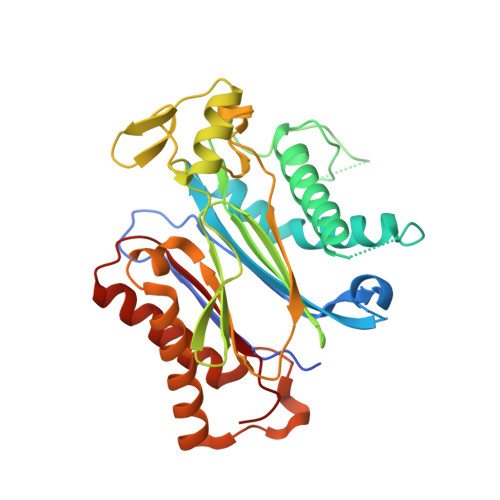
Activation of dimeric ABA receptors elicits guard cell closure, ABA-regulated gene expression, and drought tolerance.
Okamoto, M., Peterson, F.C., Defries, A., Park, S.Y., Endo, A., Nambara, E., Volkman, B.F., Cutler, S.R.(2013) Proc Natl Acad Sci U S A 110: 12132-12137
- PubMed: 23818638
- DOI: https://doi.org/10.1073/pnas.1305919110
- Primary Citation of Related Structures:
4LA7 - PubMed Abstract:
Abscisic acid (ABA) is an essential molecule in plant abiotic stress responses. It binds to soluble pyrabactin resistance1/PYR1-like/regulatory component of ABA receptor receptors and stabilizes them in a conformation that inhibits clade A type II C protein phosphatases; this leads to downstream SnRK2 kinase activation and numerous cellular outputs. We previously described the synthetic naphthalene sulfonamide ABA agonist pyrabactin, which activates seed ABA responses but fails to trigger substantial responses in vegetative tissues in Arabidopsis thaliana. Here we describe quinabactin, a sulfonamide ABA agonist that preferentially activates dimeric ABA receptors and possesses ABA-like potency in vivo. In Arabidopsis, the transcriptional responses induced by quinabactin are highly correlated with those induced by ABA treatments. Quinabactin treatments elicit guard cell closure, suppress water loss, and promote drought tolerance in adult Arabidopsis and soybean plants. The effects of quinabactin are sufficiently similar to those of ABA that it is able to rescue multiple phenotypes observed in the ABA-deficient mutant aba2. Genetic analyses show that quinabactin's effects in vegetative tissues are primarily mediated by dimeric ABA receptors. A PYL2-quinabactin-HAB1 X-ray crystal structure solved at 1.98-Å resolution shows that quinabactin forms a hydrogen bond with the receptor/PP2C "lock" hydrogen bond network, a structural feature absent in pyrabactin-receptor/PP2C complexes. Our results demonstrate that ABA receptors can be chemically controlled to enable plant protection against water stress and define the dimeric receptors as key targets for chemical modulation of vegetative ABA responses.
Organizational Affiliation:
Department of Botany and Plant Sciences and Center for Plant Cell Biology, University of California, Riverside, CA 92521, USA.