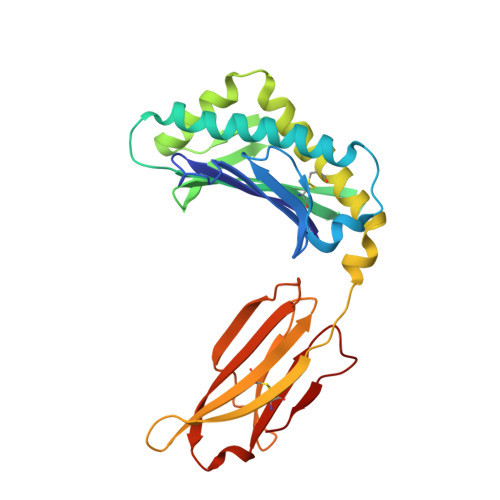
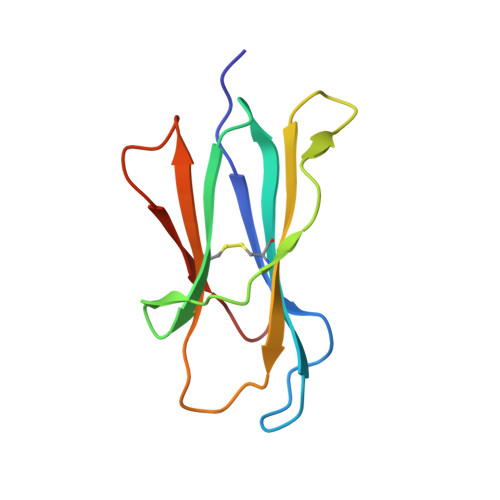
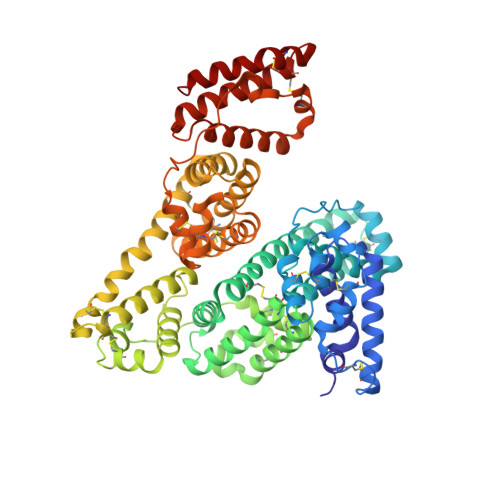
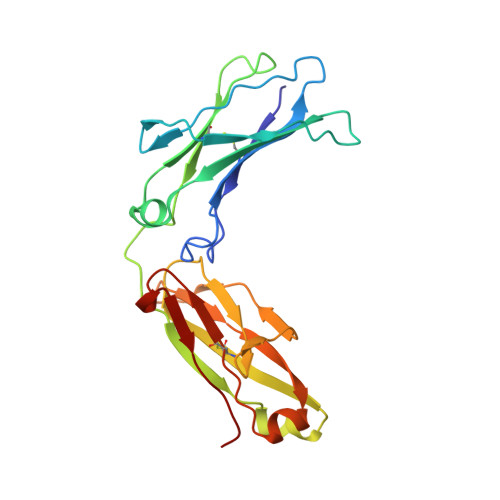
Structural Insights into Neonatal Fc Receptor-based Recycling Mechanisms.
Oganesyan, V., Damschroder, M.M., Cook, K.E., Li, Q., Gao, C., Wu, H., Dall'acqua, W.F.(2014) J Biol Chem 289: 7812-7824
- PubMed: 24469444
- DOI: https://doi.org/10.1074/jbc.M113.537563
- Primary Citation of Related Structures:
4N0F, 4N0U - PubMed Abstract:
We report the three-dimensional structure of human neonatal Fc receptor (FcRn) bound concurrently to its two known ligands. More particularly, we solved the crystal structure of the complex between human FcRn, wild-type human serum albumin (HSA), and a human Fc engineered for improved pharmacokinetics properties (Fc-YTE). The crystal structure of human FcRn bound to wild-type HSA alone is also presented. HSA domain III exhibits an extensive interface of contact with FcRn, whereas domain I plays a lesser role. A molecular explanation for the HSA recycling mechanism is provided with the identification of FcRn His(161) as the only potential direct contributor to the corresponding pH-dependent process. At last, this study also allows an accurate structural definition of residues considered for decades as important to the human IgG/FcRn interaction and reveals Fc His(310) as a significant contributor to pH-dependent binding. Finally, we explain various structural mechanisms by which several Fc mutations (including YTE) result in increased human IgG binding to FcRn. Our study provides an unprecedented relevant understanding of the molecular basis of human Fc interaction with human FcRn.
Organizational Affiliation:
From the Department of Antibody Discovery and Protein Engineering, MedImmune, Gaithersburg, Maryland 20878.