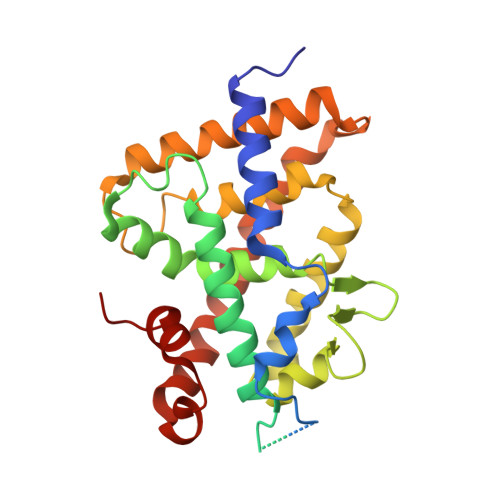
A vitamin D receptor selectively activated by gemini analogs reveals ligand dependent and independent effects.
Huet, T., Laverny, G., Ciesielski, F., Molnar, F., Ramamoorthy, T.G., Belorusova, A.Y., Antony, P., Potier, N., Metzger, D., Moras, D., Rochel, N.(2015) Cell Rep 10: 516-526
- PubMed: 25620699
- DOI: https://doi.org/10.1016/j.celrep.2014.12.045
- Primary Citation of Related Structures:
4RUJ, 4RUO, 4RUP - PubMed Abstract:
The bioactive form of vitamin D [1,25(OH)2D3] regulates mineral and bone homeostasis and exerts potent anti-inflammatory and antiproliferative properties through binding to the vitamin D receptor (VDR). The 3D structures of the VDR ligand-binding domain with 1,25(OH)2D3 or gemini analogs unveiled the molecular mechanism underlying ligand recognition. On the basis of structure-function correlations, we generated a point-mutated VDR (VDR(gem)) that is unresponsive to 1,25(OH)2D3, but the activity of which is efficiently induced by the gemini ligands. Moreover, we show that many VDR target genes are repressed by unliganded VDR(gem) and that mineral ion and bone homeostasis are more impaired in VDR(gem) mice than in VDR null mice, demonstrating that mutations abolishing VDR ligand binding result in more severe skeletal defects than VDR null mutations. As gemini ligands induce VDR(gem) transcriptional activity in mice and normalize their serum calcium levels, VDR(gem) is a powerful tool to further unravel both liganded and unliganded VDR signaling.
Organizational Affiliation:
Department of Integrative Structural Biology, Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC), Institut National de la Santé et de la Recherche Médicale (INSERM) U964, Centre National de la Recherche Scientifique (CNRS) UMR 7104, Université de Strasbourg, 67404 Illkirch, France.