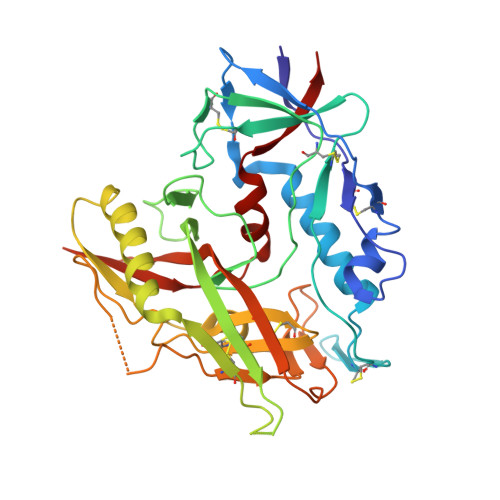
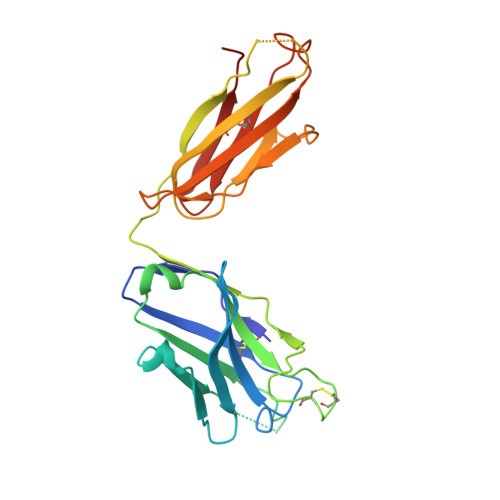
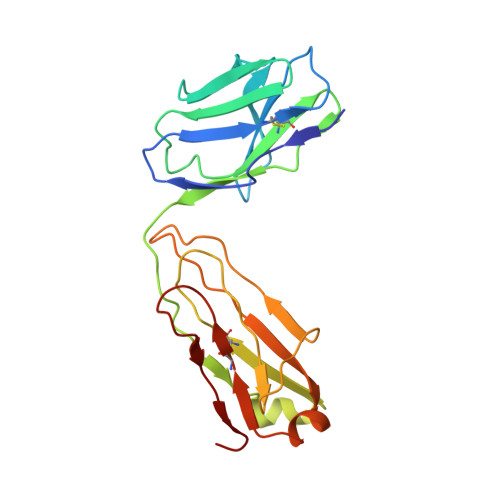
Maturation and Diversity of the VRC01-Antibody Lineage over 15 Years of Chronic HIV-1 Infection.
Wu, X., Zhang, Z., Schramm, C.A., Joyce, M.G., Do Kwon, Y., Zhou, T., Sheng, Z., Zhang, B., O'Dell, S., McKee, K., Georgiev, I.S., Chuang, G.Y., Longo, N.S., Lynch, R.M., Saunders, K.O., Soto, C., Srivatsan, S., Yang, Y., Bailer, R.T., Louder, M.K., Mullikin, J.C., Connors, M., Kwong, P.D., Mascola, J.R., Shapiro, L.(2015) Cell 161: 470-485
- PubMed: 25865483
- DOI: https://doi.org/10.1016/j.cell.2015.03.004
- Primary Citation of Related Structures:
4S1Q, 4S1R, 4S1S, 4XMP, 4XNY, 4XNZ, 4XVS, 4XVT - PubMed Abstract:
HIV-1-neutralizing antibodies develop in most HIV-1-infected individuals, although highly effective antibodies are generally observed only after years of chronic infection. Here, we characterize the rate of maturation and extent of diversity for the lineage that produced the broadly neutralizing antibody VRC01 through longitudinal sampling of peripheral B cell transcripts over 15 years and co-crystal structures of lineage members. Next-generation sequencing identified VRC01-lineage transcripts, which encompassed diverse antibodies organized into distinct phylogenetic clades. Prevalent clades maintained characteristic features of antigen recognition, though each evolved binding loops and disulfides that formed distinct recognition surfaces. Over the course of the study period, VRC01-lineage clades showed continuous evolution, with rates of ∼2 substitutions per 100 nucleotides per year, comparable to that of HIV-1 evolution. This high rate of antibody evolution provides a mechanism by which antibody lineages can achieve extraordinary diversity and, over years of chronic infection, develop effective HIV-1 neutralization.
- Vaccine Research Center, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, MD 20892, USA; Aaron Diamond AIDS Research Center, Rockefeller University, New York, NY 10016, USA.
Organizational Affiliation: