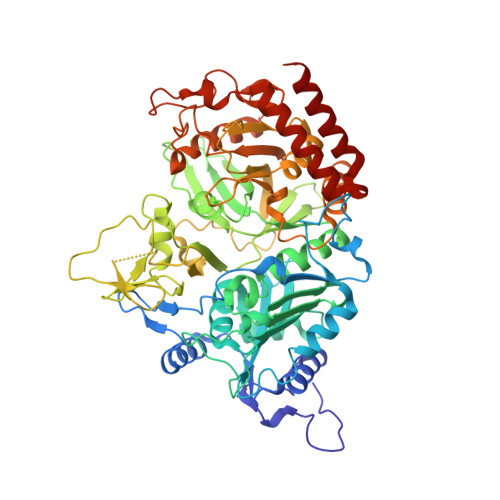
Inhibition and Allosteric Regulation of Monomeric Phosphoenolpyruvate Carboxykinase by 3-Mercaptopicolinic Acid.
Balan, M.D., Mcleod, M.J., Lotosky, W.R., Ghaly, M., Holyoak, T.(2015) Biochemistry 54: 5878-5887
- PubMed: 26322521
- DOI: https://doi.org/10.1021/acs.biochem.5b00822
- Primary Citation of Related Structures:
4YW8, 4YW9, 4YWB, 4YWD - PubMed Abstract:
For almost 40 years, it has been known that tryptophan metabolites and picolinic acid analogues act as inhibitors of gluconeogenesis. Early studies observed that 3-mercaptopicolinic acid (MPA) was a potent hypoglycemic agent via inhibition of glucose synthesis through the specific inhibition of phosphoenolpyruvate carboxykinase (PEPCK) in the gluconeogenesis pathway. Despite prior kinetic investigation, the mechanism of the inhibition by MPA is unclear. To clarify the mechanism of inhibition exerted by MPA on PEPCK, we have undertaken structural and kinetic studies. The kinetic data in concert with crystallographic structures of PEPCK in complex with MPA and the substrates for the reaction illustrate that PEPCK is inhibited by the binding of MPA at two discrete binding sites: one acting in a competitive fashion with PEP/OAA (∼10 μM) and the other acting at a previously unidentified allosteric site (Ki ∼ 150 μM). The structural studies suggest that binding of MPA to the allosteric pocket stabilizes an altered conformation of the nucleotide-binding site that in turn reduces the affinity of the enzyme for the nucleotide.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, The University of Kansas Medical Center , Kansas City, Kansas 66160, United States.