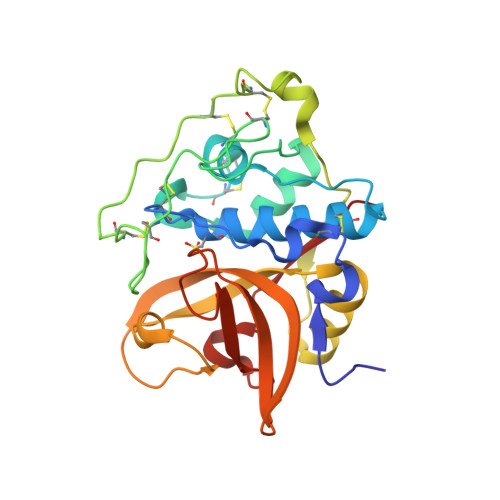
Activation route of the Schistosoma mansoni cathepsin B1 drug target: structural map with a glycosaminoglycan switch
Jilkova, A., Horn, M., Rezacova, P., Maresova, L., Fajtova, P., Brynda, J., Vondrasek, J., McKerrow, J.H., Caffrey, C.R., Mares, M.(2014) Structure 22: 1786-1798
- PubMed: 25456815
- DOI: https://doi.org/10.1016/j.str.2014.09.015
- Primary Citation of Related Structures:
4I04, 4I05, 4I07 - PubMed Abstract:
Cathepsin B1 (SmCB1) is a digestive protease of the parasitic blood fluke Schistosoma mansoni and a drug target for the treatment of schistosomiasis, a disease that afflicts over 200 million people. SmCB1 is synthesized as an inactive zymogen in which the N-terminal propeptide blocks the active site. We investigated the activation of the zymogen by which the propeptide is proteolytically removed and its regulation by sulfated polysaccharides (SPs). We determined crystal structures of three molecular forms of SmCB1 along the activation pathway: the zymogen, an activation intermediate with a partially cleaved propeptide, and the mature enzyme. We demonstrate that SPs are essential for the autocatalytic activation of SmCB1, as they interact with a specific heparin-binding domain in the propeptide. An alternative activation route is mediated by an S. mansoni asparaginyl endopeptidase (legumain) which is downregulated by SPs, indicating that SPs act as a molecular switch between both activation mechanisms.
- Institute of Organic Chemistry and Biochemistry, Academy of Sciences of the Czech Republic, 16610 Prague, Czech Republic; Department of Biochemistry, Faculty of Science, Charles University in Prague, 12843 Prague, Czech Republic.
Organizational Affiliation: