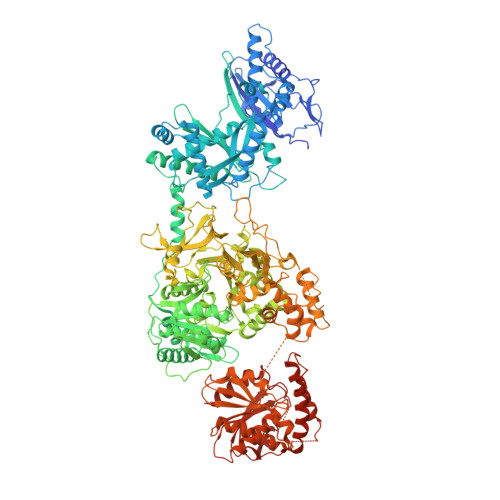
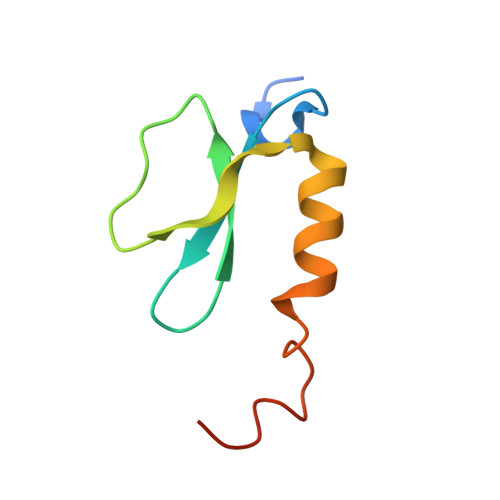
Structures of a Nonribosomal Peptide Synthetase Module Bound to MbtH-like Proteins Support a Highly Dynamic Domain Architecture.
Miller, B.R., Drake, E.J., Shi, C., Aldrich, C.C., Gulick, A.M.(2016) J Biol Chem 291: 22559-22571
- PubMed: 27597544
- DOI: https://doi.org/10.1074/jbc.M116.746297
- Primary Citation of Related Structures:
5JA1, 5JA2 - PubMed Abstract:
Nonribosomal peptide synthetases (NRPSs) produce a wide variety of peptide natural products. During synthesis, the multidomain NRPSs act as an assembly line, passing the growing product from one module to the next. Each module generally consists of an integrated peptidyl carrier protein, an amino acid-loading adenylation domain, and a condensation domain that catalyzes peptide bond formation. Some adenylation domains interact with small partner proteins called MbtH-like proteins (MLPs) that enhance solubility or activity. A structure of an MLP bound to an adenylation domain has been previously reported using a truncated adenylation domain, precluding any insight that might be derived from understanding the influence of the MLP on the intact adenylation domain or on the dynamics of the entire NRPS module. Here, we present the structures of the full-length NRPS EntF bound to the MLPs from Escherichia coli and Pseudomonas aeruginosa These new structures, along with biochemical and bioinformatics support, further elaborate the residues that define the MLP-adenylation domain interface. Additionally, the structures highlight the dynamic behavior of NRPS modules, including the module core formed by the adenylation and condensation domains as well as the orientation of the mobile thioesterase domain.
Organizational Affiliation:
From the Hauptman-Woodward Medical Research Institute, Buffalo, New York 14203.