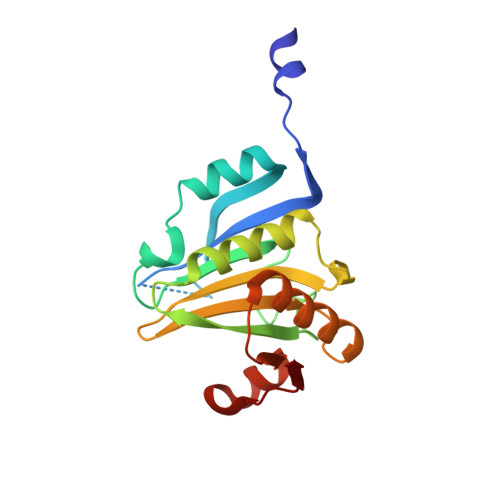
Structure of eIF4E in Complex with an eIF4G Peptide Supports a Universal Bipartite Binding Mode for Protein Translation.
Miras, M., Truniger, V., Silva, C., Verdaguer, N., Aranda, M.A., Querol-Audi, J.(2017) Plant Physiol 174: 1476-1491
- PubMed: 28522457
- DOI: https://doi.org/10.1104/pp.17.00193
- Primary Citation of Related Structures:
5ME5, 5ME6, 5ME7 - PubMed Abstract:
The association-dissociation of the cap-binding protein eukaryotic translation initiation factor 4E (eIF4E) with eIF4G is a key control step in eukaryotic translation. The paradigm on the eIF4E-eIF4G interaction states that eIF4G binds to the dorsal surface of eIF4E through a single canonical alpha-helical motif, while metazoan eIF4E-binding proteins (m4E-BPs) advantageously compete against eIF4G via bimodal interactions involving this canonical motif and a second noncanonical motif of the eIF4E surface. Metazoan eIF4Gs share this extended binding interface with m4E-BPs, with significant implications on the understanding of translation regulation and the design of therapeutic molecules. Here we show the high-resolution structure of melon ( Cucumis melo ) eIF4E in complex with a melon eIF4G peptide and propose the first eIF4E-eIF4G structural model for plants. Our structural data together with functional analyses demonstrate that plant eIF4G binds to eIF4E through both the canonical and noncanonical motifs, similarly to metazoan eIF4E-eIF4G complexes. As in the case of metazoan eIF4E-eIF4G, this may have very important practical implications, as plant eIF4E-eIF4G is also involved in a significant number of plant diseases. In light of our results, a universal eukaryotic bipartite mode of binding to eIF4E is proposed.
- Centro de Edafología y Biología Aplicada del Segura (CEBAS), Consejo Superior de Investigaciones Científicas (CSIC), 30100 Espinardo, Murcia, Spain.
Organizational Affiliation: