The evolutionary advantage of an aromatic clamp in plant family 3 glycoside exo-hydrolases.
Luang, S., Fernandez-Luengo, X., Nin-Hill, A., Streltsov, V.A., Schwerdt, J.G., Alonso-Gil, S., Ketudat Cairns, J.R., Pradeau, S., Fort, S., Marechal, J.D., Masgrau, L., Rovira, C., Hrmova, M.(2022) Nat Commun 13: 5577-5577
- PubMed: 36151080
- DOI: https://doi.org/10.1038/s41467-022-33180-5
- Primary Citation of Related Structures:
6JG1, 6JG2, 6JG6, 6JG7, 6JGA, 6JGB, 6JGC, 6JGD, 6JGE, 6JGG, 6JGK, 6JGL, 6JGN, 6JGO, 6JGP, 6JGQ, 6JGR, 6JGS, 6JGT, 6K6V, 6KUF, 6L1J, 6LBB, 6LBV, 6LC5 - PubMed Abstract:
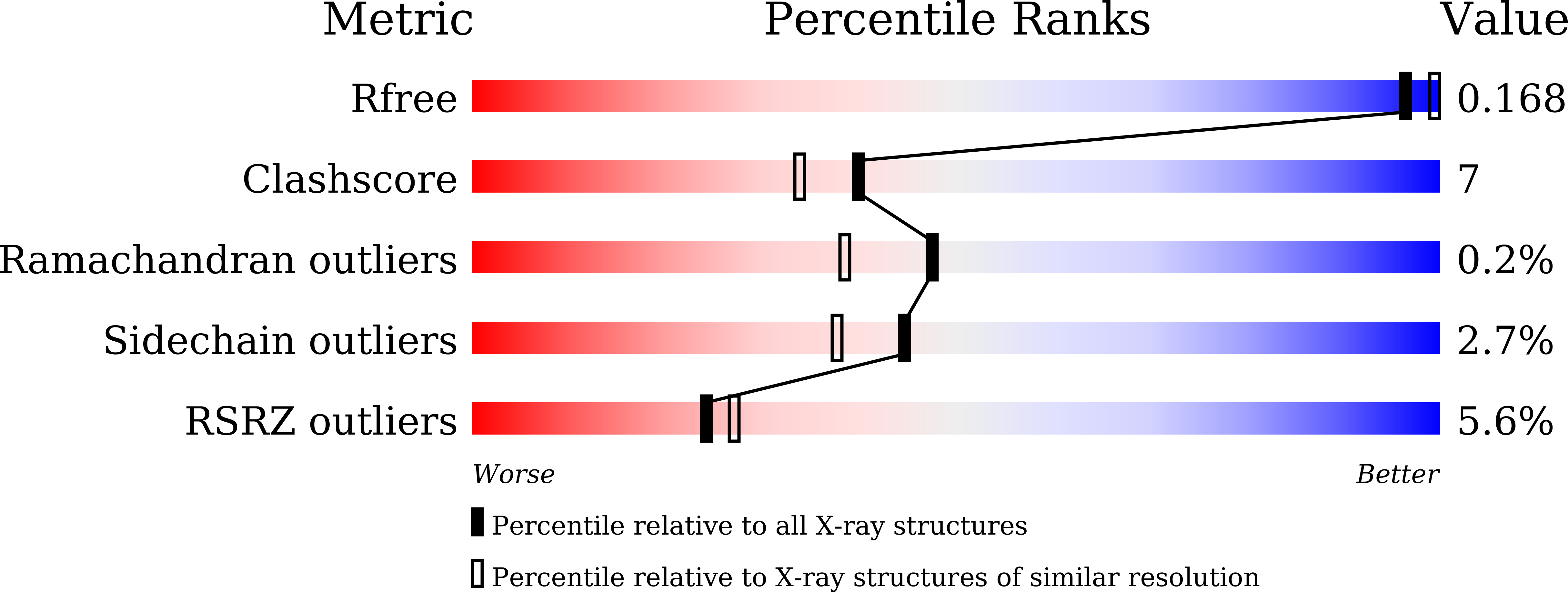
In the barley β-D-glucan glucohydrolase, a glycoside hydrolase family 3 (GH3) enzyme, the Trp286/Trp434 clamp ensures β-D-glucosides binding, which is fundamental for substrate hydrolysis during plant growth and development. We employ mutagenesis, high-resolution X-ray crystallography, and multi-scale molecular modelling methods to examine the binding and conformational behaviour of isomeric β-D-glucosides during substrate-product assisted processive catalysis that operates in GH3 hydrolases. Enzyme kinetics reveals that the W434H mutant retains broad specificity, while W434A behaves as a strict (1,3)-β-D-glucosidase. Investigations of reactant movements on the nanoscale reveal that processivity is sensitive to mutation-specific alterations of the tryptophan clamp. While wild-type and W434H utilise a lateral cavity for glucose displacement and sliding of (1,3)-linked hydrolytic products through the catalytic site without dissociation, consistent with their high hydrolytic rates, W434A does not adopt processive catalysis. Phylogenomic analyses of GH3 hydrolases disclose the evolutionary advantage of the tryptophan clamp that confers broad specificity, high catalytic efficiency, and processivity.
Organizational Affiliation:
School of Agriculture, Food and Wine, and Waite Research Institute, University of Adelaide, Waite Research Precinct, Glen Osmond, SA, Australia.