Atypical TRAV1-2 - T cell receptor recognition of the antigen-presenting molecule MR1.
Awad, W., Meermeier, E.W., Sandoval-Romero, M.L., Le Nours, J., Worley, A.H., Null, M.D., Liu, L., McCluskey, J., Fairlie, D.P., Lewinsohn, D.M., Rossjohn, J.(2020) J Biological Chem 295: 14445-14457
- PubMed: 32817339
- DOI: https://doi.org/10.1074/jbc.RA120.015292
- Primary Citation of Related Structures:
6XQP, 6XQQ - PubMed Abstract:
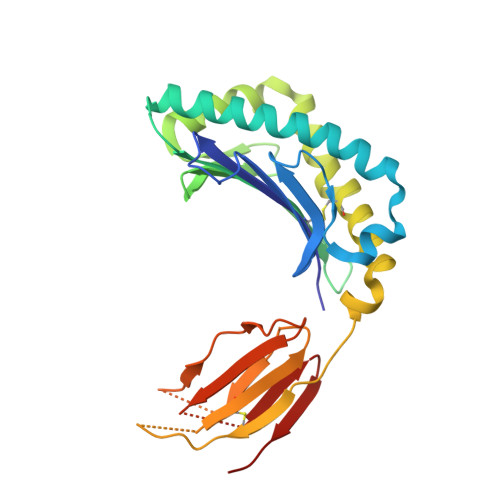
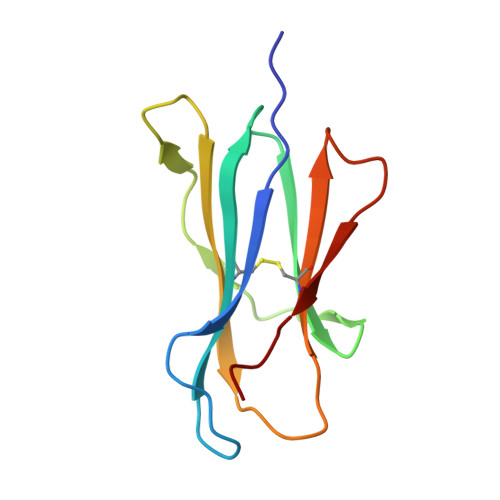
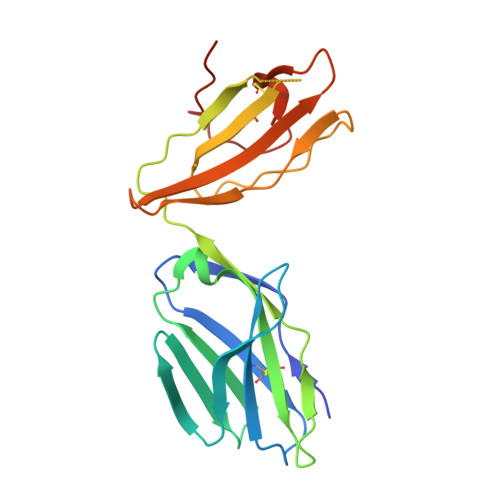
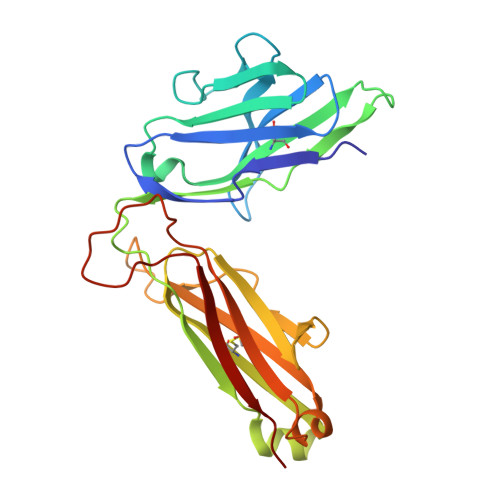
MR1 presents vitamin B-related metabolites to mucosal associated invariant T (MAIT) cells, which are characterized, in part, by the TRAV1-2 + αβ T cell receptor (TCR). In addition, a more diverse TRAV1-2 - MR1-restricted T cell repertoire exists that can possess altered specificity for MR1 antigens. However, the molecular basis of how such TRAV1-2 - TCRs interact with MR1-antigen complexes remains unclear. Here, we describe how a TRAV12-2 + TCR (termed D462-E4) recognizes an MR1-antigen complex. We report the crystal structures of the unliganded D462-E4 TCR and its complex with MR1 presenting the riboflavin-based antigen 5-OP-RU. Here, the TRBV29-1 β-chain of the D462-E4 TCR binds over the F'-pocket of MR1, whereby the complementarity-determining region (CDR) 3β loop surrounded and projected into the F'-pocket. Nevertheless, the CDR3β loop anchored proximal to the MR1 A'-pocket and mediated direct contact with the 5-OP-RU antigen. The D462-E4 TCR footprint on MR1 contrasted that of the TRAV1-2 + and TRAV36 + TCRs' docking topologies on MR1. Accordingly, diverse MR1-restricted T cell repertoire reveals differential docking modalities on MR1, thus providing greater scope for differing antigen specificities.
- Infection and Immunity Program and Department of Biochemistry and Molecular Biology, Biomedicine Discovery Institute, Monash University, Clayton, Victoria, Australia.
Organizational Affiliation: