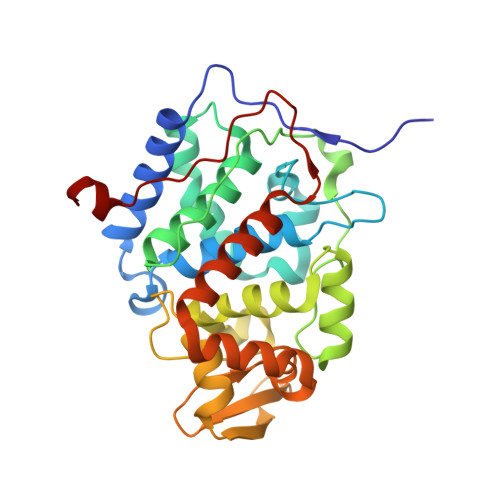
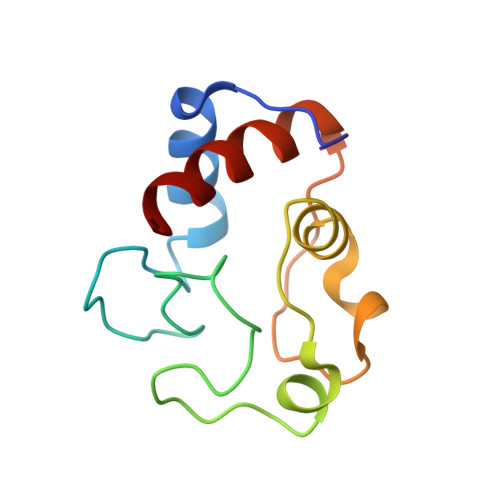
Tuning Radical Relay Residues by Proton Management Rescues Protein Electron Hopping.
Yee, E.F., Dzikovski, B., Crane, B.R.(2019) J Am Chem Soc 141: 17571-17587
- PubMed: 31603693
- DOI: https://doi.org/10.1021/jacs.9b05715
- Primary Citation of Related Structures:
6P41, 6P42, 6P43 - PubMed Abstract:
Transient tyrosine and tryptophan radicals play key roles in the electron transfer (ET) reactions of photosystem (PS) II, ribonucleotide reductase (RNR), photolyase, and many other proteins. However, Tyr and Trp are not functionally interchangeable, and the factors controlling their reactivity are often unclear. Cytochrome c peroxidase (CcP) employs a Trp191 •+ radical to oxidize reduced cytochrome c ( Cc ). Although a Tyr191 replacement also forms a stable radical, it does not support rapid ET from Cc . Here we probe the redox properties of CcP Y191 by non-natural amino acid substitution, altering the ET driving force and manipulating the protic environment of Y191. Higher potential fluorotyrosine residues increase ET rates marginally, but only addition of a hydrogen bond donor to Tyr191 • (via Leu232His or Glu) substantially alters activity by increasing the ET rate by nearly 30-fold. ESR and ESEEM spectroscopies, crystallography, and pH-dependent ET kinetics provide strong evidence for hydrogen bond formation to Y191 • by His232/Glu232. Rate measurements and rapid freeze quench ESR spectroscopy further reveal differences in radical propagation and Cc oxidation that support an increased Y191 • formal potential of ∼200 mV in the presence of E232. Hence, Y191 inactivity results from a potential drop owing to Y191 •+ deprotonation. Incorporation of a well-positioned base to accept and donate back a hydrogen bond upshifts the Tyr • potential into a range where it can effectively oxidize Cc . These findings have implications for the Y Z /Y D radicals of PS II, hole-hopping in RNR and cryptochrome, and engineering proteins for long-range ET reactions.
Organizational Affiliation:
Department of Chemistry and Chemical Biology , Cornell University , Ithaca , New York 14853 , United States.