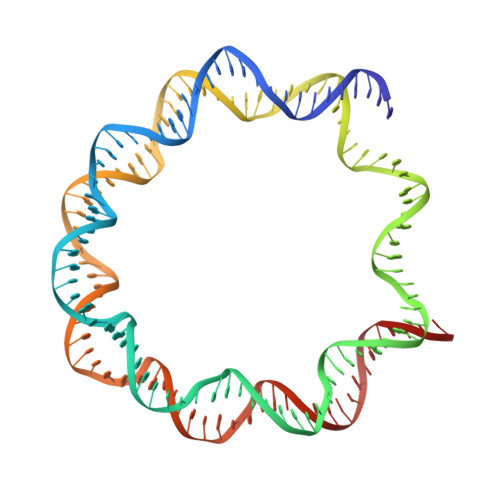
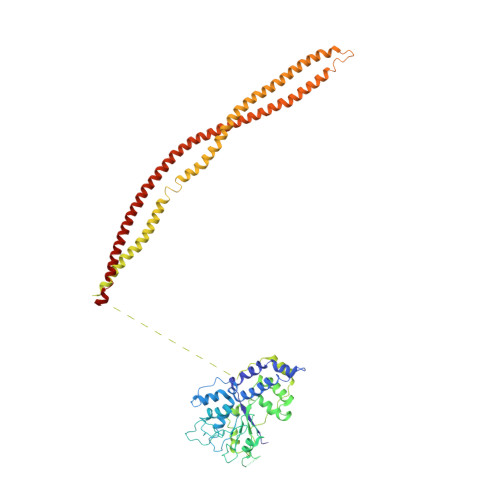
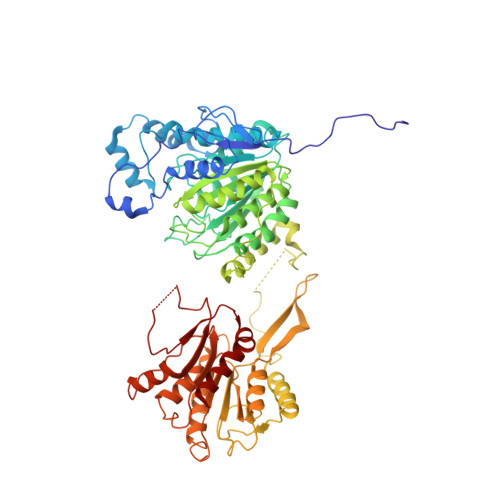
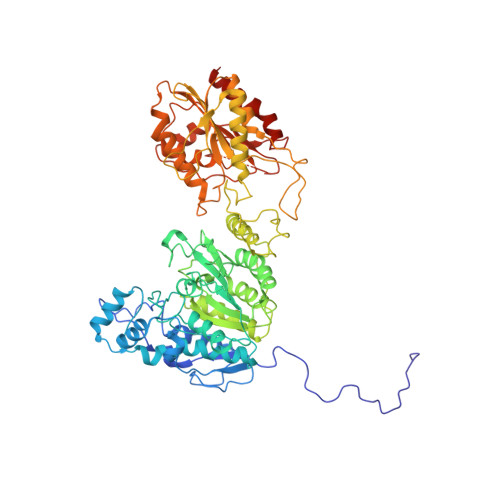
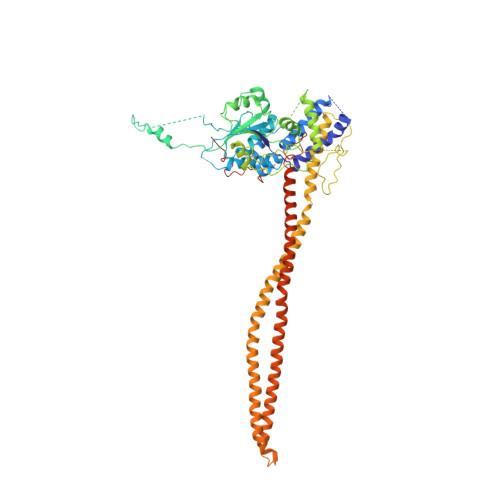
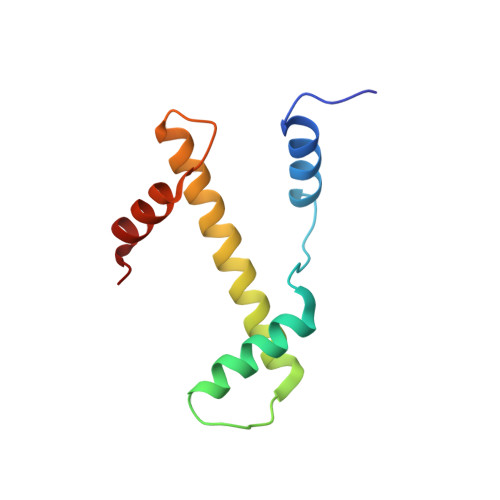
Structural basis for the regulation of nucleosome recognition and HDAC activity by histone deacetylase assemblies.
Lee, J.H., Bollschweiler, D., Schafer, T., Huber, R.(2021) Sci Adv 7
- PubMed: 33523989
- DOI: https://doi.org/10.1126/sciadv.abd4413
- Primary Citation of Related Structures:
6Z6F, 6Z6H, 6Z6O, 6Z6P - PubMed Abstract:
The chromatin-modifying histone deacetylases (HDACs) remove acetyl groups from acetyl-lysine residues in histone amino-terminal tails, thereby mediating transcriptional repression. Structural makeup and mechanisms by which multisubunit HDAC complexes recognize nucleosomes remain elusive. Our cryo-electron microscopy structures of the yeast class II HDAC ensembles show that the HDAC protomer comprises a triangle-shaped assembly of stoichiometry Hda1 2 -Hda2-Hda3, in which the active sites of the Hda1 dimer are freely accessible. We also observe a tetramer of protomers, where the nucleosome binding modules are inaccessible. Structural analysis of the nucleosome-bound complexes indicates how positioning of Hda1 adjacent to histone H2B affords HDAC catalysis. Moreover, it reveals how an intricate network of multiple contacts between a dimer of protomers and the nucleosome creates a platform for expansion of the HDAC activities. Our study provides comprehensive insight into the structural plasticity of the HDAC complex and its functional mechanism of chromatin modification.
- Max Planck Institute of Biochemistry, Am Klopferspitz 18, 82152 Martinsried, Germany. hlee@biochem.mpg.de huber@biochem.mpg.de.
Organizational Affiliation: