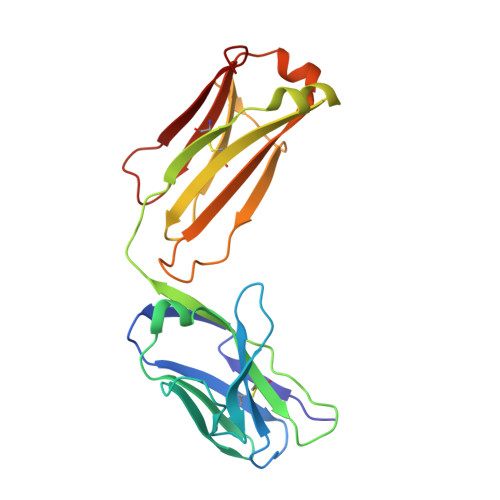
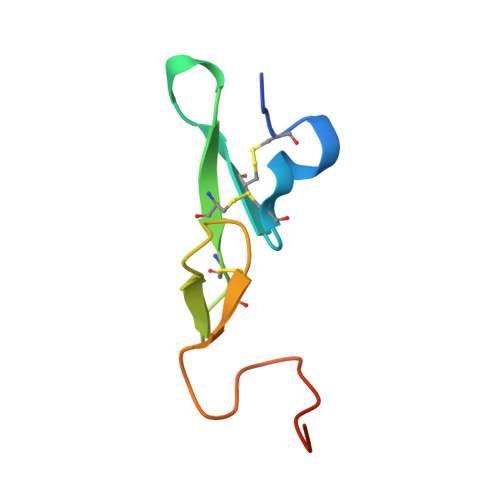
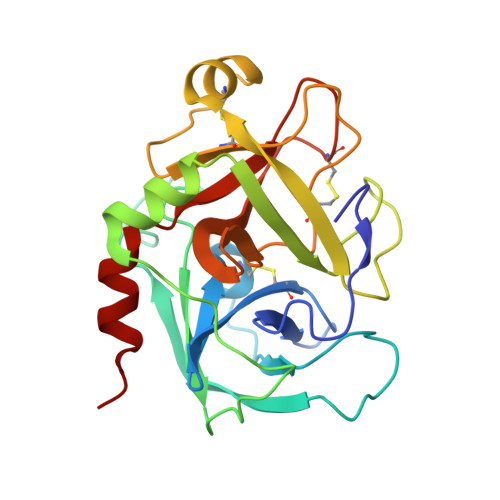
A factor VIIIa-mimetic bispecific antibody, Mim8, ameliorates bleeding upon severe vascular challenge in hemophilia A mice.
Ostergaard, H., Lund, J., Greisen, P.J., Kjellev, S., Henriksen, A., Lorenzen, N., Johansson, E., Roder, G., Rasch, M.G., Johnsen, L.B., Egebjerg, T., Lund, S., Rahbek-Nielsen, H., Gandhi, P.S., Lamberth, K., Loftager, M., Andersen, L.M., Bonde, A.C., Stavenuiter, F., Madsen, D.E., Li, X., Holm, T.L., Ley, C.D., Thygesen, P., Zhu, H., Zhou, R., Thorn, K., Yang, Z., Hermit, M.B., Bjelke, J.R., Hansen, B.G., Hilden, I.(2021) Blood 138: 1258-1268
- PubMed: 34077951
- DOI: https://doi.org/10.1182/blood.2020010331
- Primary Citation of Related Structures:
7AHU, 7AHV - PubMed Abstract:
Hemophilia A is a bleeding disorder resulting from deficient factor VIII (FVIII), which normally functions as a cofactor to activated factor IX (FIXa) that facilitates activation of factor X (FX). To mimic this property in a bispecific antibody format, a screening was conducted to identify functional pairs of anti-FIXa and anti-FX antibodies, followed by optimization of functional and biophysical properties. The resulting bispecific antibody (Mim8) assembled efficiently with FIXa and FX on membranes, and supported activation with an apparent equilibrium dissociation constant of 16 nM. Binding affinity with FIXa and FX in solution was much lower, with equilibrium dissociation constant values for FIXa and FX of 2.3 and 1.5 µM, respectively. In addition, the activity of Mim8 was dependent on stimulatory activity contributed by the anti-FIXa arm, which enhanced the proteolytic activity of FIXa by 4 orders of magnitude. In hemophilia A plasma and whole blood, Mim8 normalized thrombin generation and clot formation, with potencies 13 and 18 times higher than a sequence-identical analogue of emicizumab. A similar potency difference was observed in a tail vein transection model in hemophilia A mice, whereas reduction of bleeding in a severe tail-clip model was observed only for Mim8. Furthermore, the pharmacokinetic parameters of Mim8 were investigated and a half-life of 14 days shown in cynomolgus monkeys. In conclusion, Mim8 is an activated FVIII mimetic with a potent and efficacious hemostatic effect based on preclinical data.
- Global Drug Discovery and.
Organizational Affiliation: