A crystallography-based study of fragment extensions into the 14-3-3 binding groove
Centorrino, F., Wu, Q., Cossar, P., Brunsveld, L., Ottmann, C.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| 14-3-3 protein sigma | 253 | Homo sapiens | Mutation(s): 0 Gene Names: SFN, HME1 | 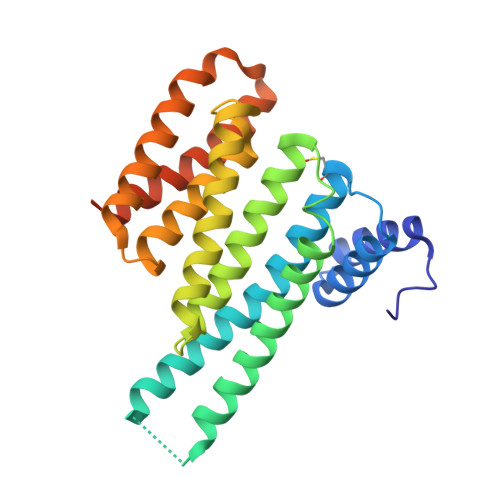 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P31947 (Homo sapiens) Explore P31947 Go to UniProtKB: P31947 | |||||
PHAROS: P31947 GTEx: ENSG00000175793 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P31947 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Find similar proteins by: Sequence | 3D Structure
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Neurogenic locus notch homolog protein 4 | 11 | Homo sapiens | Mutation(s): 0 | 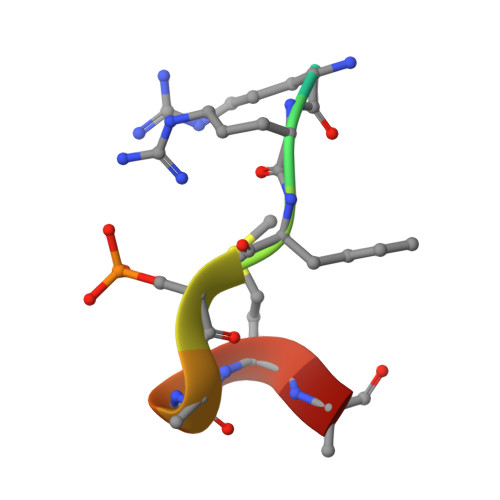 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for Q99466 (Homo sapiens) Explore Q99466 Go to UniProtKB: Q99466 | |||||
PHAROS: Q99466 GTEx: ENSG00000204301 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q99466 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| 0B7 (Subject of Investigation/LOI) Query on 0B7 | E [auth A] | ~{N}-[(5-carbamimidoyl-3-phenyl-thiophen-2-yl)methyl]-1~{H}-indole-6-carboxamide C21 H18 N4 O S AWDSOCIOIXYPGZ-UHFFFAOYSA-N |  | ||
| CL Query on CL | C [auth A] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| MG Query on MG | D [auth A] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Modified Residues 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| CSO Query on CSO | A | L-PEPTIDE LINKING | C3 H7 N O3 S |  | CYS |
| SEP Query on SEP | B | L-PEPTIDE LINKING | C3 H8 N O6 P |  | SER |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 82.437 | α = 90 |
| b = 111.431 | β = 90 |
| c = 62.424 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| DIALS | data reduction |
| Aimless | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| H2020 Marie Curie Actions of the European Commission | European Union | 675179 |